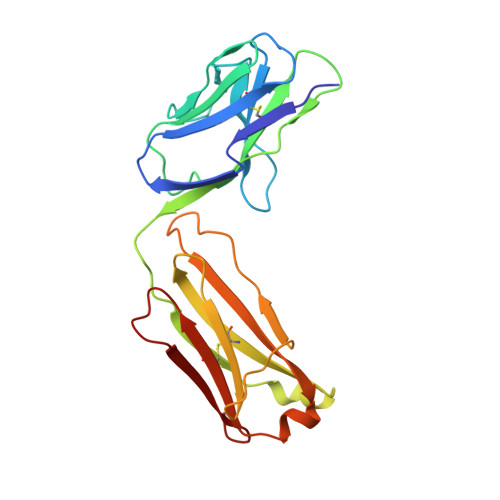
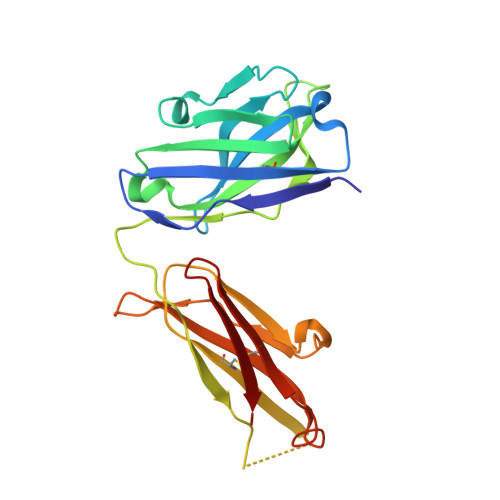
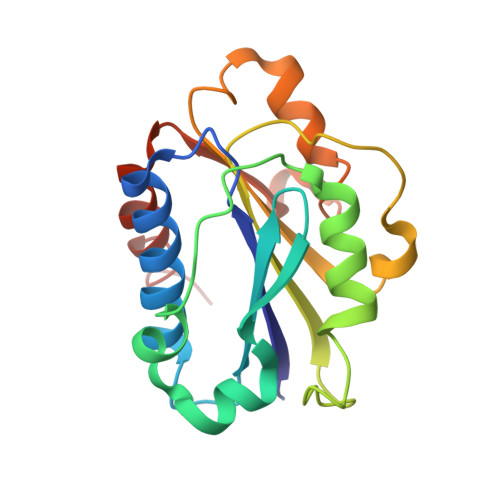
Efalizumab binding to the LFA-1 alphaL I domain blocks ICAM-1 binding via steric hindrance.
Li, S., Wang, H., Peng, B., Zhang, M., Zhang, D., Hou, S., Guo, Y., Ding, J.(2009) Proc Natl Acad Sci U S A 106: 4349-4354
- PubMed: 19258452 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.0810844106
- Primary Citation Related Structures:
3EO9, 3EOA, 3EOB - PubMed Abstract:
Lymphocyte function-associated antigen 1 (LFA-1) plays important roles in immune cell adhesion, trafficking, and activation and is a therapeutic target for the treatment of multiple autoimmune diseases. Efalizumab is one of the most efficacious antibody drugs for treating psoriasis, a very common skin disease, through inhibition of the binding of LFA-1 to the ligand intercellular adhesion molecule 1 (ICAM-1). We report here the crystal structures of the Efalizumab Fab alone and in complex with the LFA-1 alpha(L) I domain, which reveal the molecular mechanism of inhibition of LFA-1 by Efalizumab. The Fab binds with an epitope on the inserted (I) domain that is distinct from the ligand-binding site. Efalizumab binding blocks the binding of LFA-1 to ICAM-1 via steric hindrance between its light chain and ICAM-1 domain 2 and thus inhibits the activities of LFA-1. These results have important implications for the development of improved antibodies and new therapeutic strategies for the treatment of autoimmune diseases.
- State Key Laboratory of Molecular Biology and Research Center for Structural Biology, Institute of Biochemistry and Cell Biology, Chinese Academy of Sciences, 320 Yue-Yang Road, Shanghai 200031, China.
Organizational Affiliation: