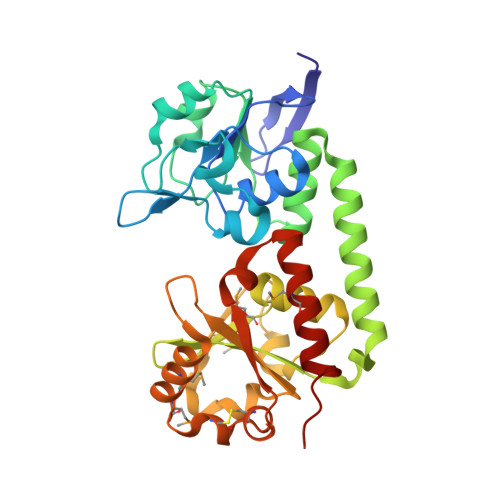
Trapping open and closed forms of FitE-A group III periplasmic binding protein.
Shi, R., Proteau, A., Wagner, J., Cui, Q., Purisima, E.O., Matte, A., Cygler, M.(2008) Proteins 75: 598-609
- PubMed: 19004000 Search on PubMed
- DOI: https://doi.org/10.1002/prot.22272
- Primary Citation Related Structures:
3BE5, 3BE6 - PubMed Abstract:
Periplasmic binding proteins (PBPs) are essential components of bacterial transport systems, necessary for bacterial growth and survival. The two-domain structures of PBPs are topologically classified into three groups based on the number of crossovers or hinges between the globular domains: group I PBPs have three connections, group II have two, and group III have only one. Although a large number of structures for group I or II PBPs are known, fewer group III PBPs have been structurally characterized. Group I and II PBPs exhibit significant domain motions during transition from the unbound to ligand-bound form, however, no large conformational changes have been observed to date in group III PBPs. We have solved the crystal structure of a periplasmic binding protein FitE, part of an iron transport system, fit, recently identified in a clinical E. coli isolate. The structure, determined at 1.8 A resolution, shows that FitE is a group III PBP containing a single alpha-helix bridging the two domains. Among the individual FitE molecules present in two crystal forms we observed three different conformations (open, closed, intermediate). Our crystallographic and molecular dynamics results strongly support the notion that group III PBPs also adopt the same Venus flytrap mechanism as do groups I and II PBPs. Unlike other group III PBPs, FitE forms dimers both in solution and in the crystals. The putative siderophore binding pocket is lined with arginine residues, suggesting an anionic nature of the iron-containing siderophore.
- Department of Biochemistry, McGill University, Montréal, Québec, Canada.
Organizational Affiliation: