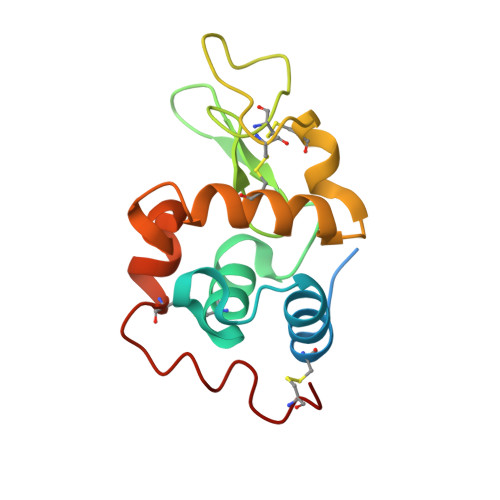
Protecting role of cosolvents in protein denaturation by SDS: a structural study.
Michaux, C., Pouyez, J., Wouters, J., Prive, G.G.(2008) BMC Struct Biol 8: 29-35
- PubMed: 18522744 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1186/1472-6807-8-29
- Primary Citation Related Structures:
3B6L, 3B72 - PubMed Abstract:
Recently, we reported a unique approach to preserve the activity of some proteins in the presence of the denaturing agent, Sodium Dodecyl Sulfate (SDS). This was made possible by addition of the amphipathic solvent 2,4-Methyl-2-PentaneDiol (MPD), used as protecting but also as refolding agent for these proteins. Although the persistence of the protein activity in the SDS/MPD mixture was clearly established, preservation of their structure was only speculative until now. In this paper, a detailed X-ray study addresses the pending question. Crystals of hen egg-white lysozyme were grown for the first time in the presence of MPD and denaturing concentrations of SDS. Depending on crystallization conditions, tetragonal crystals in complex with either SDS or MPD were collected. The conformation of both structures was very similar to the native lysozyme and the obtained complexes of SDS-lysozyme and MPD-lysozyme give some insights in the interplay of protein-SDS and protein-MPD interactions. This study clearly established the preservation of the enzyme structure in a SDS/MPD mixture. It is hypothesized that high concentrations of MPD would change the properties of SDS and lower or avoid interactions between the denaturant and the protein. These structural data therefore support the hypothesis that MPD avoids disruption of the enzyme structure by SDS and can protect proteins from SDS denaturation.
- Chemistry department, CBS lab, CPTS group, 61 rue de Bruxelles, B-5000 Namur, Belgium. catherine.michaux@fundp.ac.be
Organizational Affiliation: