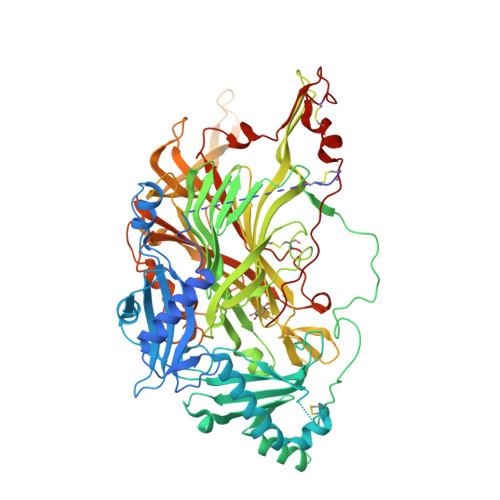
A new crystal form of human vascular adhesion protein 1
Ernberg, K., McGrath, A.P., Peat, T.S., Adams, T.E., Xiao, X., Pham, T., Newman, J., McDonald, I.A., Collyer, C.A., Guss, J.M.(2010) Acta Crystallogr Sect F Struct Biol Cryst Commun 66: 1572-1578
- PubMed: 21139198 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S1744309110041515
- Primary Citation Related Structures:
3ALA - PubMed Abstract:
Human vascular adhesion protein 1 (VAP-1) is involved in lymphocyte-endothelial cell adhesion and has been implicated in many human inflammatory diseases. VAP-1 is a member of the copper amine oxidase family of enzymes with a trihydroxyphenylalanine quinone (TPQ) cofactor. Previously characterized crystals of VAP-1 suffered from anisotropy and contained disordered regions; in addition, one form was consistently twinned. In an effort to grow crystals that diffracted to higher resolution for inhibitor-binding studies, a construct with an N-terminal deletion was made and expressed in the Chinese hamster ovary (CHO) glycosylation mutant cell line Lec8. Screening produced crystals that displayed some anisotropy and contained seven molecules per asymmetric unit. These crystals belonged to space group C2, with unit-cell parameters a=394.5, b=115.8, c=179.3 Å, β=112.3°. The structure was refined to a resolution of 2.9 Å, with Rcryst and Rfree values of 0.250 and 0.286, respectively.
- School of Molecular Bioscience, University of Sydney, NSW 2006, Australia.
Organizational Affiliation: