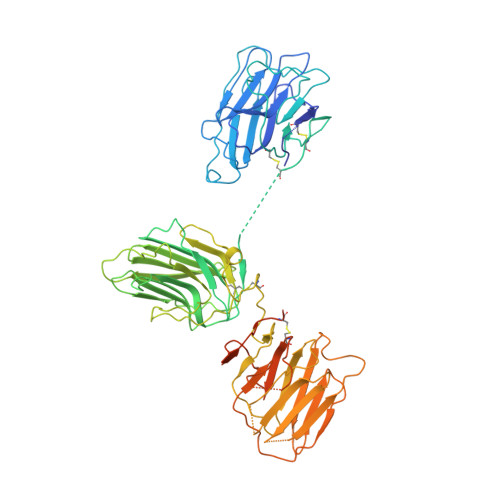
Crystal Structure of the Lg1-3 Region of the Laminin {Alpha}2 Chain.
Carafoli, F., Clout, N.J., Hohenester, E.(2009) J Biological Chem 284: 22786
- PubMed: 19553699 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M109.026658
- Primary Citation Related Structures:
2WJS - PubMed Abstract:
Laminins are large heterotrimeric glycoproteins with many essential functions in basement membrane assembly and function. Cell adhesion to laminins is mediated by a tandem of five laminin G-like (LG) domains at the C terminus of the alpha chain. Integrin binding requires an intact LG1-3 region, as well as contributions from the coiled coil formed by the alpha, beta, and gamma chains. We have determined the crystal structure at 2.8-A resolution of the LG1-3 region of the laminin alpha2 chain (alpha 2LG1-3). The three LG domains adopt typical beta-sandwich folds, with canonical calcium binding sites in LG1 and LG2. LG2 and LG3 interact through a substantial interface, but LG1 is completely dissociated from the LG2-3 pair. We suggest that the missing gamma chain tail may be required to stabilize the interaction between LG1 and LG2-3 in the biologically active conformation. A global analysis of N-linked glycosylation sites shows that the beta-sandwich faces of LG1 are free of carbohydrate modifications in all five laminin alpha chains, suggesting that these surfaces may harbor the integrin binding site. The alpha 2LG1-3 structure provides the first atomic view of the integrin binding region of laminins.
- Department of Life Sciences, Imperial College London, London SW7 2AZ, United Kingdom.
Organizational Affiliation: