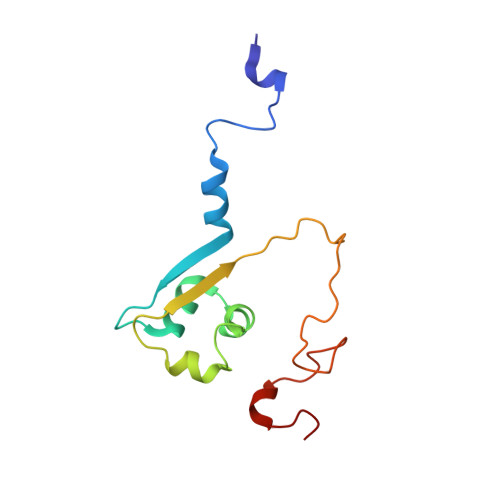
The Intimin periplasmic domain mediates dimerisation and binding to peptidoglycan.
Leo, J.C., Oberhettinger, P., Chaubey, M., Schutz, M., Kuhner, D., Bertsche, U., Schwarz, H., Gotz, F., Autenrieth, I.B., Coles, M., Linke, D.(2015) Mol Microbiol 95: 80-100
- PubMed: 25353290 Search on PubMed
- DOI: https://doi.org/10.1111/mmi.12840
- Primary Citation Related Structures:
2MPW - PubMed Abstract:
Intimin and Invasin are prototypical inverse (Type Ve) autotransporters and important virulence factors of enteropathogenic Escherichia coli and Yersinia spp. respectively. In addition to a C-terminal extracellular domain and a β-barrel transmembrane domain, both proteins also contain a short N-terminal periplasmic domain that, in Intimin, includes a lysin motif (LysM), which is thought to mediate binding to peptidoglycan. We show that the periplasmic domain of Intimin does bind to peptidoglycan both in vitro and in vivo, but only under acidic conditions. We were able to determine a dissociation constant of 0.8 μM for this interaction, whereas the Invasin periplasmic domain, which lacks a LysM, bound only weakly in vitro and failed to bind peptidoglycan in vivo. We present the solution structure of the Intimin LysM, which has an additional α-helix conserved within inverse autotransporter LysMs but lacking in others. In contrast to previous reports, we demonstrate that the periplasmic domain of Intimin mediates dimerisation. We further show that dimerisation and peptidoglycan binding are general features of LysM-containing inverse autotransporters. Peptidoglycan binding by the periplasmic domain in the infection process may aid in resisting mechanical and chemical stress during transit through the gastrointestinal tract.
- Department of Protein Evolution, Max Planck Institute for Developmental Biology, 72076, Tübingen, Germany.
Organizational Affiliation: