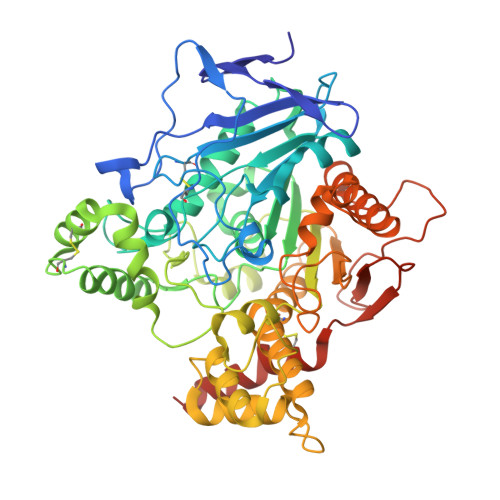
Crystal structure of thioflavin T bound to the peripheral site of Torpedo californica acetylcholinesterase reveals how thioflavin T acts as a sensitive fluorescent reporter of ligand binding to the acylation site.
Harel, M., Sonoda, L.K., Silman, I., Sussman, J.L., Rosenberry, T.L.(2008) J Am Chem Soc 130: 7856-7861
- PubMed: 18512913 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/ja7109822
- Primary Citation Related Structures:
2J3Q - PubMed Abstract:
Acetylcholinesterase plays a key role in cholinergic synaptic transmission by hydrolyzing the neurotransmitter acetylcholine with one of the highest known catalytic rate constants. Hydrolysis occurs in a narrow and deep gorge that contains two sites of ligand binding: A peripheral site, or P-site, near the gorge entrance that contributes to catalytic efficiency both by transiently trapping substrate molecules as they enter the gorge and by allosterically accelerating the transfer of the substrate acyl group to a serine hydroxyl in an acylation site or A-site at the base of the gorge. Thioflavin T is a useful reporter of ligand interactions with the A-site. It binds specifically to the P-site with fluorescence that is enhanced approximately 1000-fold over that of unbound thioflavin T, and the enhanced fluorescence is quenched 1.5- to 4-fold when another ligand binds to the A-site in a ternary complex. To clarify the structural basis of this advantageous signal change, we here report the X-ray structure of the complex of thioflavin T with Torpedo californica acetylcholinesterase. The two aromatic rings in thioflavin T are coplanar and are packed snugly parallel to the aromatic side chains of Trp279, Tyr334, and Phe330. Overlays of this structure with the crystal structures of Torpedo californica acetylcholinesterase complexes with either edrophonium or m-( N, N, N-trimethylammonio)-2,2,2-trifluoroacetophenone, two small aromatic ligands that bind specifically to the A-site, indicate that the phenyl side chain of Phe330 must rotate to sterically accommodate both thioflavin T and the A-site ligand in the ternary complex. This rotation may allow some relaxation of the strict coplanarity of the aromatic rings in the bound thioflavin T and result in partial quenching of its fluorescence.
- Department of Structural Biology, Weizmann Institute of Science, Rehovot 76100, Israel.
Organizational Affiliation: