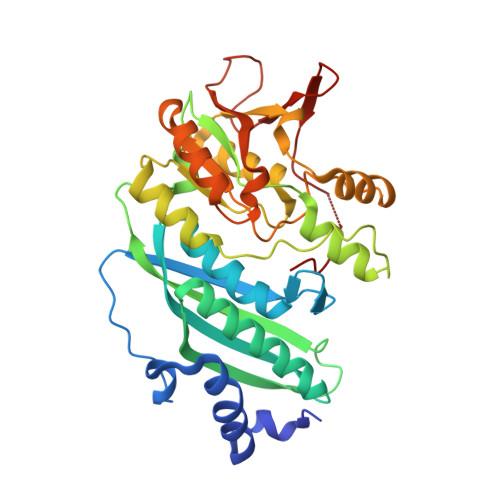
Crystal Structures of Hydrogenase Maturation Protein HypE in the Apo and ATP-bound Forms
Shomura, Y., Komori, H., Miyabe, N., Tomiyama, M., Shibata, N., Higuchi, Y.(2007) J Mol Biology 372: 1045-1054
- PubMed: 17706667 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2007.07.023
- Primary Citation Related Structures:
2Z1T, 2Z1U - PubMed Abstract:
The hydrogenase maturation protein HypE serves an essential function in the biosynthesis of the nitrile group, which is subsequently coordinated to Fe as CN(-) ligands in [Ni-Fe] hydrogenase. Here, we present the crystal structures of HypE from Desulfovibrio vulgaris Hildenborough in the presence and in the absence of ATP at a resolution of 2.0 A and 2.6 A, respectively. Comparison of the apo structure with the ATP-bound structure reveals that binding ATP causes an induced-fit movement of the N-terminal portion, but does not entail an overall structural change. The residue Cys341 at the C terminus, whose thiol group is supposed to be carbamoylated before the nitrile group synthesis, is completely buried within the protein and is located in the vicinity of the gamma-phosphate group of the bound ATP. This suggests that the catalytic reaction occurs in this configuration but that a conformational change is required for the carbamoylation of Cys341. A glutamate residue is found close to the thiol group as well, which is suggestive of deprotonation of the carbamoyl group at the beginning of the reactions.
- Department of Life Science, Graduate School of Life Science, University of Hyogo, 3-2-1 Koto, Kamigori-cho, Ako-gun, Hyogo 678-1297, Japan; RIKEN SPring-8 Center, 1-1-1 Koto, Sayo-gun, Sayo-cho, Hyogo 679-5148, Japan. Electronic address: shomura@sci.u-hyogo.ac.jp.
Organizational Affiliation: