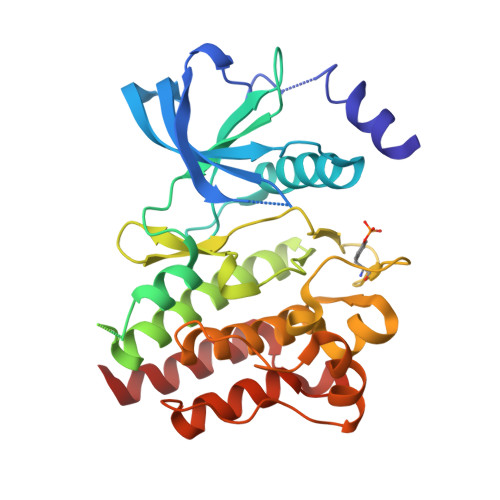
Synthesis, structure-activity relationship and crystallographic studies of 3-substituted indolin-2-one RET inhibitors.
Mologni, L., Rostagno, R., Brussolo, S., Knowles, P.P., Kjaer, S., Murray-Rust, J., Rosso, E., Zambon, A., Scapozza, L., McDonald, N.Q., Lucchini, V., Gambacorti-Passerini, C.(2010) Bioorg Med Chem 18: 1482-1496
- PubMed: 20117004
- DOI: https://doi.org/10.1016/j.bmc.2010.01.011
- Primary Citation of Related Structures:
2X2K, 2X2L, 2X2M - PubMed Abstract:
The synthesis, structure-activity relationships (SAR) and structural data of a series of indolin-2-one inhibitors of RET tyrosine kinase are described. These compounds were designed to explore the available space around the indolinone scaffold within RET active site. Several substitutions at different positions were tested and biochemical data were used to draw a molecular model of steric and electrostatic interactions, which can be applied to design more potent and selective RET inhibitors. The crystal structures of RET kinase domain in complex with three inhibitors were solved. All three compounds bound in the ATP pocket and formed two hydrogen bonds with the kinase hinge region. Crystallographic analysis confirmed predictions from molecular modelling and helped refine SAR results. These data provide important information for the development of indolinone inhibitors for the treatment of RET-driven cancers.
- Department of Clinical Medicine and Prevention, University of Milano-Bicocca, Monza, Italy. luca.mologni@unimib.it
Organizational Affiliation: