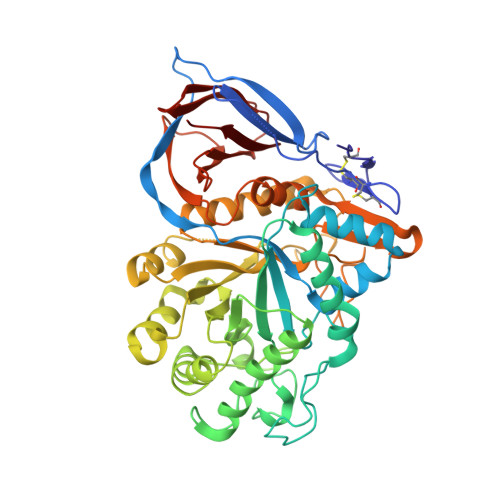
Characterization of Gene-Activated Human Acid-Beta-Glucosidase: Crystal Structure, Glycan Composition and Internalization Into Macrophages.
Brumshtein, B., Salinas, P., Peterson, B., Chan, V., Silman, I., Sussman, J.L., Savickas, P.J., Robinson, G.S., Futerman, A.H.(2010) Glycobiology 20: 24
- PubMed: 19741058 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/glycob/cwp138
- Primary Citation Related Structures:
2WKL - PubMed Abstract:
Gaucher disease, the most common lysosomal storage disease, can be treated with enzyme replacement therapy (ERT), in which defective acid-beta-glucosidase (GlcCerase) is supplemented by a recombinant, active enzyme. The X-ray structures of recombinant GlcCerase produced in Chinese hamster ovary cells (imiglucerase, Cerezyme) and in transgenic carrot cells (prGCD) have been previously solved. We now describe the structure and characteristics of a novel form of GlcCerase under investigation for the treatment of Gaucher disease, Gene-Activated human GlcCerase (velaglucerase alfa). In contrast to imiglucerase and prGCD, velaglucerase alfa contains the native human enzyme sequence. All three GlcCerases consist of three domains, with the active site located in domain III. The distances between the carboxylic oxygens of the catalytic residues, E340 and E235, are consistent with distances proposed for acid-base hydrolysis. Kinetic parameters (K(m) and V(max)) of velaglucerase alfa and imiglucerase, as well as their specific activities, are similar. However, analysis of glycosylation patterns shows that velaglucerase alfa displays distinctly different structures from imiglucerase and prGCD. The predominant glycan on velaglucerase alfa is a high-mannose type, with nine mannose units, while imiglucerase contains a chitobiose tri-mannosyl core glycan with fucosylation. These differences in glycosylation affect cellular internalization; the rate of velaglucerase alfa internalization into human macrophages is at least 2-fold greater than that of imiglucerase.
- Department of Neurobiology, Weizmann Institute of Science, Rehovot 76100, Israel.
Organizational Affiliation: