T Cell-Mediated Autoimmune Disease due to Low-Affinity Crossreactivity to Common Microbial Peptides.
Harkiolaki, M., Holmes, S.L., Svendsen, P., Gregersen, J.W., Jensen, L.T., Mcmahon, R., Friese, M.A., Van Boxel, G., Etzensperger, R., Tzartos, J.S., Kranc, K., Sainsbury, S., Harlos, K., Mellins, E.D., Palace, J., Esiri, M.M., Van Der Merwe, P.A., Jones, E.Y., Fugger, L.(2009) Immunity 30: 348
- PubMed: 19303388 Search on PubMed
- DOI: https://doi.org/10.1016/j.immuni.2009.01.009
- Primary Citation Related Structures:
2WBJ - PubMed Abstract:
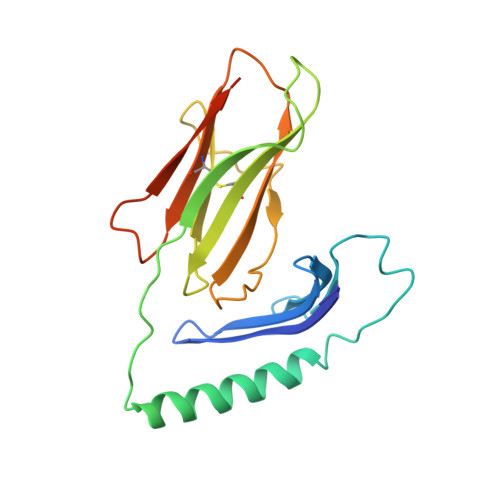
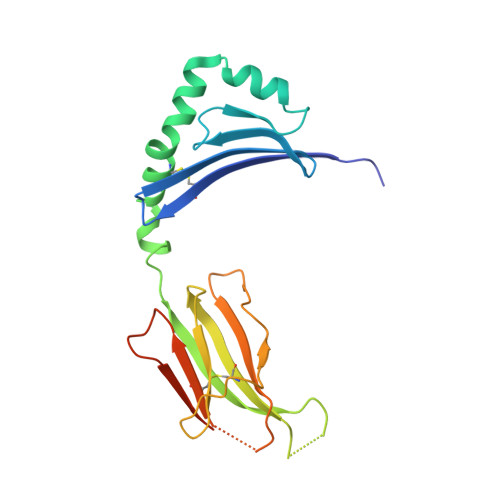
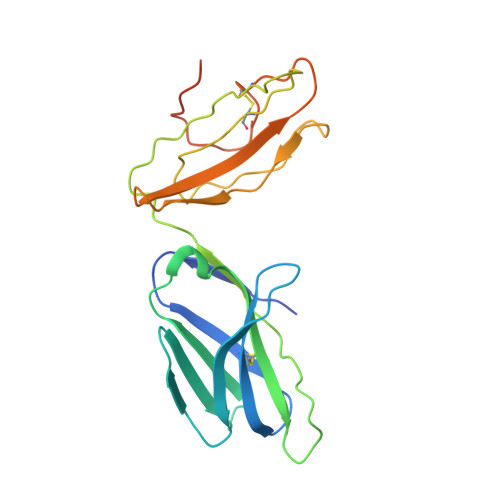
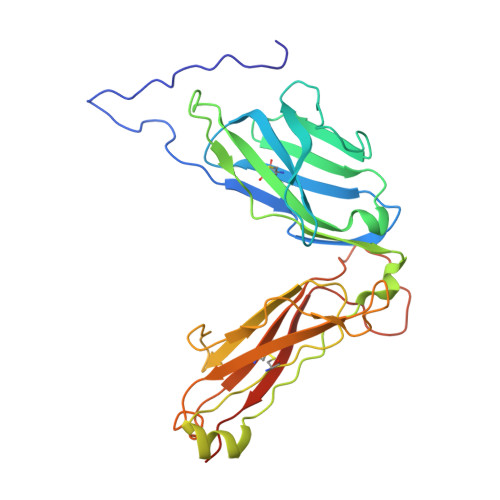
Environmental factors account for 75% of the risk of developing multiple sclerosis (MS). Numerous infections have been suspected as environmental disease triggers, but none of them has consistently been incriminated, and it is unclear how so many different infections may play a role. We show that a microbial peptide, common to several major classes of bacteria, can induce MS-like disease in humanized mice by crossreacting with a T cell receptor (TCR) that also recognizes a peptide from myelin basic protein, a candidate MS autoantigen. Structural analysis demonstrates this crossreactivity is due to structural mimicry of a binding hotspot shared by self and microbial antigens, rather than to degenerate TCR recognition. Biophysical studies reveal that the autoreactive TCR binding affinity is markedly lower for the microbial (mimicry) peptide than for the autoantigenic peptide. Thus, these data suggest a possible explanation for the difficulty in incriminating individual infections in the development of MS.
- Division of Structural Biology, Henry Wellcome Building for Genomic Medicine, University of Oxford, Roosevelt Drive, Oxford OX37BN, UK.
Organizational Affiliation: