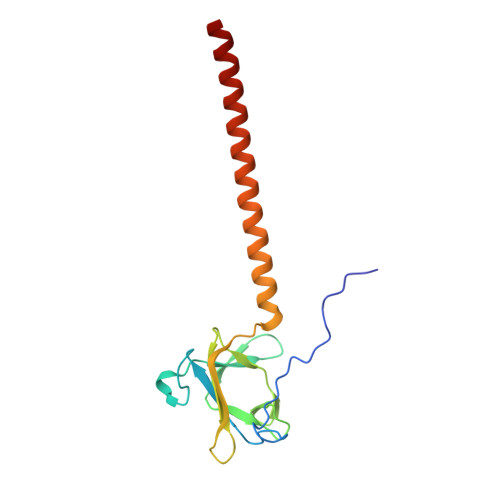
Bacteriophage P22 Tailspike: Structure of the Complete Protein and Function of the Interdomain Linker
Seul, A., Mueller, J.J., Andres, D., Stettner, E., Heinemann, U., Seckler, R.(2014) Acta Crystallogr D Biol Crystallogr 70: 1336-1345
- PubMed: 24816102 Search on PubMed
- DOI: https://doi.org/10.1107/S1399004714002685
- Primary Citation Related Structures:
2VKY, 2VNL, 2XC1 - PubMed Abstract:
Attachment of phages to host cells, followed by phage DNA ejection, represents the first stage of viral infection of bacteria. Salmonella phage P22 has been extensively studied, serving as an experimental model for bacterial infection by phages. P22 engages bacteria by binding to the sugar moiety of lipopolysaccharides using the viral tailspike protein for attachment. While the structures of the N-terminal particle-binding domain and the major receptor-binding domain of the tailspike have been analyzed individually, the three-dimensional organization of the intact protein, including the highly conserved linker region between the two domains, remained unknown. A single amino-acid exchange in the linker sequence made it possible to crystallize the full-length protein. Two crystal structures of the linker region are presented: one attached to the N-terminal domain and the other present within the complete tailspike protein. Both retain their biological function, but the mutated full-length tailspike displays a retarded folding pathway. Fitting of the full-length tailspike into a published cryo-electron microscopy map of the P22 virion requires an elastic distortion of the crystal structure. The conservation of the linker suggests a role in signal transmission from the distal tip of the molecule to the phage head, eventually leading to DNA ejection.
- Physikalische Biochemie, Universität Potsdam, Karl-Liebknecht-Strasse 24-25, 14476 Potsdam, Germany.
Organizational Affiliation: