Analysis of Nasturtium Tmnxg1 Complexes by Crystallography and Molecular Dynamics Provides Detailed Insight Into Substrate Recognition by Family Gh16 Xyloglucan Endo-Transglycosylases and Endo-Hydrolases.
Mark, P., Baumann, M.J., Eklof, J.M., Gullfot, F., Michel, G., Kallas, A.M., Teeri, T.T., Brumer, H., Czjzek, M.(2009) Proteins 75: 820
- PubMed: 19004021 Search on PubMed
- DOI: https://doi.org/10.1002/prot.22291
- Primary Citation Related Structures:
2VH9 - PubMed Abstract:
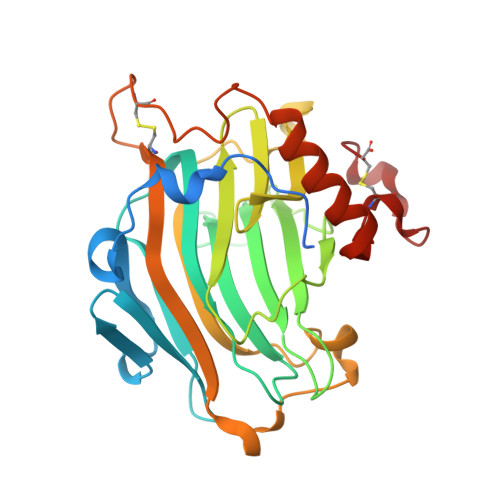
Reorganization and degradation of the wall crosslinking and seed storage polysaccharide xyloglucan by glycoside hydrolase family 16 (GH16) endo-transglycosylases and hydrolases are crucial to the growth of the majority of land plants, affecting processes as diverse as germination, morphogenesis, and fruit ripening. A high-resolution, three-dimensional structure of a nasturtium (Tropaeolum majus) endo-xyloglucanase loop mutant, TmNXG1-DeltaYNIIG, with an oligosaccharide product bound in the negative active-site subsites, has been solved by X-ray crystallography. Comparison of this novel complex to that of the strict xyloglucan endo-transglycosylase PttXET16-34 from hybrid aspen (Populus tremula x tremuloides), previously solved with a xylogluco-oligosaccharide bound in the positive subsites, highlighted key protein structures that affect the disparate catalytic activities displayed by these closely related enzymes. Combination of these "partial" active-site complexes through molecular dynamics simulations in water allowed modeling of wild-type TmNXG1, TmNXG1-DeltaYNIIG, and wild-type PttXET16-34 in complex with a xyloglucan octadecasaccharide spanning the entire catalytic cleft. A comprehensive analysis of these full-length complexes underscored the importance of various loops lining the active site. Subtle differences leading to a tighter hydrogen bonding pattern on the negative (glycosyl donor) binding subsites, together with loop flexibility on the positive (glycosyl acceptor) binding subsites appear to favor hydrolysis over transglycosylation in GH16 xyloglucan-active enzymes.
- School of Biotechnology, Royal Institute of Technology (KTH), AlbaNova University Centre. S 106 91 Stockholm, Sweden.
Organizational Affiliation: