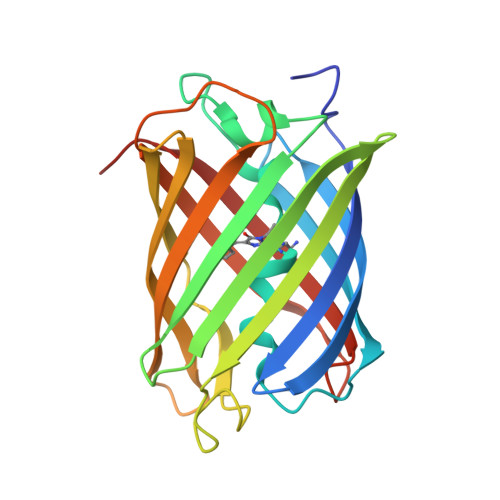
Structural Rearrangements Near the Chromophore Influence the Maturation Speed and Brightness of Dsred Variants.
Strongin, D.E., Bevis, B., Khuong, N., Downing, M.E., Strack, R.L., Sundaram, K., Glick, B.S., Keenan, R.J.(2007) Protein Eng Des Sel 20: 525
- PubMed: 17962222 Search on PubMed
- DOI: https://doi.org/10.1093/protein/gzm046
- Primary Citation Related Structures:
2VAD, 2VAE - PubMed Abstract:
The red fluorescent protein DsRed has been extensively engineered for use as an in vivo research tool. In fast maturing DsRed variants, the chromophore maturation half-time is approximately 40 min, compared to approximately 12 h for wild-type DsRed. Further, DsRed has been converted from a tetramer into a monomer, a task that entailed mutating approximately 20% of the amino acids. These engineered variants of DsRed have proven extremely valuable for biomedical research, but the structural basis for the improved characteristics has not been thoroughly investigated. Here we present a 1.7 A crystal structure of the fast maturing tetrameric variant DsRed.T4. We also present a biochemical characterization and 1.6 A crystal structure of the monomeric variant DsRed.M1, also known as DsRed-Monomer. Analysis of the crystal structures suggests that rearrangements of Ser69 and Glu215 contribute to fast maturation, and that positioning of the Lys70 side chain modulates fluorescence quantum yield. Despite the 45 mutations in DsRed.M1 relative to wild-type DsRed, there is a root-mean-square deviation of only 0.3 A between the two structures. We propose that novel intramolecular interactions in DsRed.M1 partially compensate for the loss of intermolecular interactions found in the tetramer.
- Department of Molecular Genetics and Cell Biology, University of Chicago, IL 60637, USA.
Organizational Affiliation: