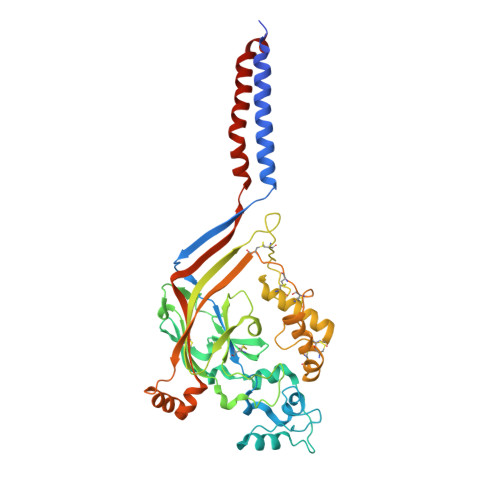
Structure of acid-sensing ion channel 1 at 1.9A resolution and low pH
Jasti, J., Furukawa, H., Gonzales, E.B., Gouaux, E.(2007) Nature 449: 316-323
- PubMed: 17882215 Search on PubMed
- DOI: https://doi.org/10.1038/nature06163
- Primary Citation Related Structures:
2QTS - PubMed Abstract:
Acid-sensing ion channels (ASICs) are voltage-independent, proton-activated receptors that belong to the epithelial sodium channel/degenerin family of ion channels and are implicated in perception of pain, ischaemic stroke, mechanosensation, learning and memory. Here we report the low-pH crystal structure of a chicken ASIC1 deletion mutant at 1.9 A resolution. Each subunit of the chalice-shaped homotrimer is composed of short amino and carboxy termini, two transmembrane helices, a bound chloride ion and a disulphide-rich, multidomain extracellular region enriched in acidic residues and carboxyl-carboxylate pairs within 3 A, suggesting that at least one carboxyl group bears a proton. Electrophysiological studies on aspartate-to-asparagine mutants confirm that these carboxyl-carboxylate pairs participate in proton sensing. Between the acidic residues and the transmembrane pore lies a disulphide-rich 'thumb' domain poised to couple the binding of protons to the opening of the ion channel, thus demonstrating that proton activation involves long-range conformational changes.
- Vollum Institute, Oregon Health and Science University, 3181 SW Sam Jackson Park Road, Portland, Oregon 97239, USA.
Organizational Affiliation: