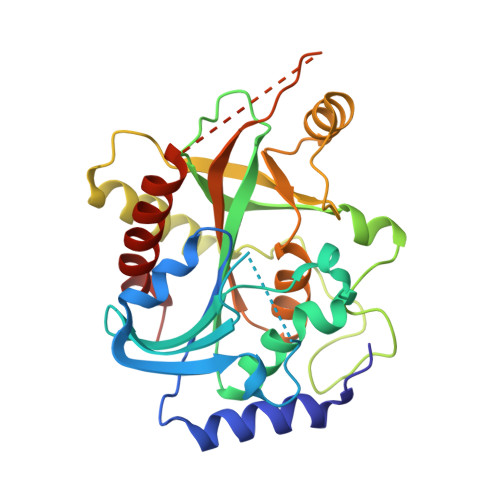
Crystal structure of calf spleen purine nucleoside phosphorylase complexed to a novel purine analogue.
Pereira, H.M., Berdini, V., Cleasby, A., Garratt, R.C.(2007) FEBS Lett 581: 5082-5086
- PubMed: 17927987 Search on PubMed
- DOI: https://doi.org/10.1016/j.febslet.2007.09.051
- Primary Citation Related Structures:
2QPL - PubMed Abstract:
The combined use of a rapid virtual screen of a small fragment library together with a single point enzyme assay has been used for the discovery of novel PNP inhibitors. The availability of readily soakable crystals of bovine PNP has allowed the approach to be experimentally validated by determining the crystal structure of one of the inhibitor-PNP complexes. Comparison of the experimentally determined binding mode with that predicted by the virtual screening shows them to be similar. This represents a starting point for the growth of the ligand into a higher affinity inhibitor.
- Instituto de Física de São Carlos, Universidade de São Paulo, Physics and Informatics, Av. Trabalhador Saocarlense 400, Centro CP 369, 13560-970 Sao Carlos, São Paulo, Brazil. hmuniz.pereira@gmail.com
Organizational Affiliation: