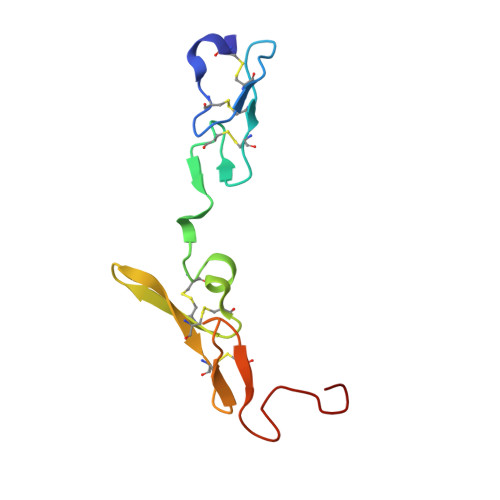
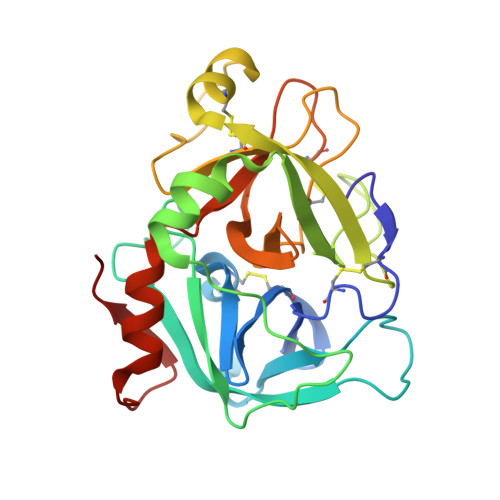
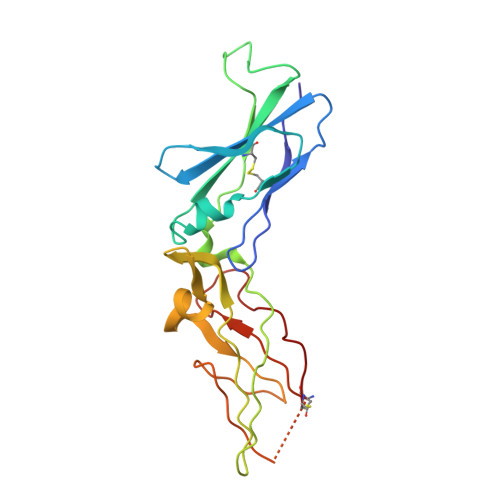
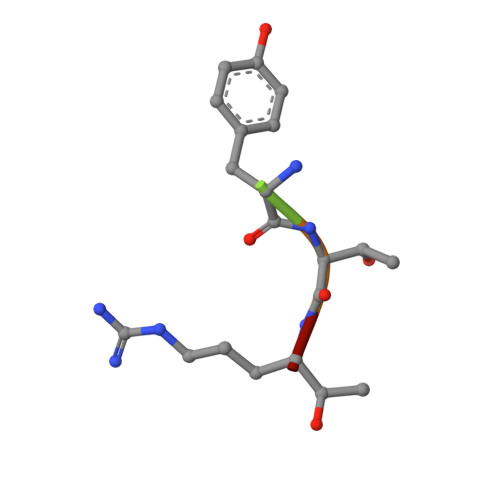
Engineering the substrate and inhibitor specificities of human coagulation Factor VIIa
Larsen, K.S., Ostergaard, H., Bjelke, J.R., Olsen, O.H., Rasmussen, H.B., Christensen, L., Kragelund, B.B., Stennicke, H.R.(2007) Biochem J 405: 429-438
- PubMed: 17456045 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1042/BJ20061901
- Primary Citation Related Structures:
2PUQ - PubMed Abstract:
The remarkably high specificity of the coagulation proteases towards macromolecular substrates is provided by numerous interactions involving the catalytic groove and remote exosites. For FVIIa [activated FVII (Factor VII)], the principal initiator of coagulation via the extrinsic pathway, several exosites have been identified, whereas only little is known about the specificity dictated by the active-site architecture. In the present study, we have profiled the primary P4-P1 substrate specificity of FVIIa using positional scanning substrate combinatorial libraries and evaluated the role of the selective active site in defining specificity. Being a trypsin-like serine protease, FVIIa had P1 specificity exclusively towards arginine and lysine residues. In the S2 pocket, threonine, leucine, phenylalanine and valine residues were the most preferred amino acids. Both S3 and S4 appeared to be rather promiscuous, however, with some preference for aromatic amino acids at both positions. Interestingly, a significant degree of interdependence between the S3 and S4 was observed and, as a consequence, the optimal substrate for FVIIa could not be derived directly from a subsite-directed specificity screen. To evaluate the role of the active-site residues in defining specificity, a series of mutants of FVIIa were prepared at position 239 (position 99 in chymotrypsin), which is considered to be one of the most important residues for determining P2 specificity of the trypsin family members. This was confirmed for FVIIa by marked changes in primary substrate specificity and decreased rates of antithrombin III inhibition. Interestingly, these changes do not necessarily coincide with an altered ability to activate Factor X, demonstrating that inhibitor and macromolecular substrate selectivity may be engineered separately.
- Haemostasis Biochemistry, Novo Nordisk A/S, Novo Nordisk Park, DK-2760 Måløv, Denmark. ksln@novonordisk.com
Organizational Affiliation: