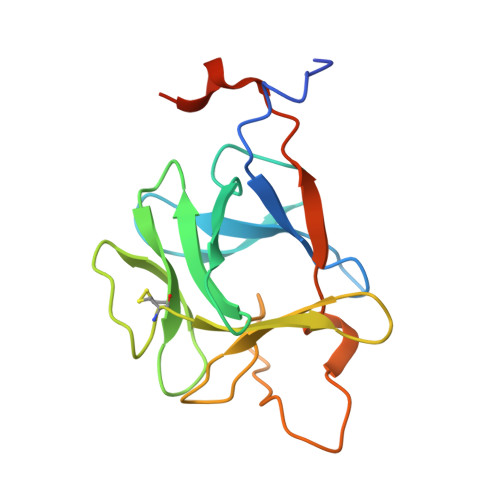
Molecular insights into the klotho-dependent, endocrine mode of action of fibroblast growth factor 19 subfamily members.
Goetz, R., Beenken, A., Ibrahimi, O.A., Kalinina, J., Olsen, S.K., Eliseenkova, A.V., Xu, C., Neubert, T.A., Zhang, F., Linhardt, R.J., Yu, X., White, K.E., Inagaki, T., Kliewer, S.A., Yamamoto, M., Kurosu, H., Ogawa, Y., Kuro-O, M., Lanske, B., Razzaque, M.S., Mohammadi, M.(2007) Mol Cell Biol 27: 3417-3428
- PubMed: 17339340 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1128/MCB.02249-06
- Primary Citation Related Structures:
2P23, 2P39 - PubMed Abstract:
Unique among fibroblast growth factors (FGFs), FGF19, -21, and -23 act in an endocrine fashion to regulate energy, bile acid, glucose, lipid, phosphate, and vitamin D homeostasis. These FGFs require the presence of Klotho/betaKlotho in their target tissues. Here, we present the crystal structures of FGF19 alone and FGF23 in complex with sucrose octasulfate, a disaccharide chemically related to heparin. The conformation of the heparin-binding region between beta strands 10 and 12 in FGF19 and FGF23 diverges completely from the common conformation adopted by paracrine-acting FGFs. A cleft between this region and the beta1-beta2 loop, the other heparin-binding region, precludes direct interaction between heparin/heparan sulfate and backbone atoms of FGF19/23. This reduces the heparin-binding affinity of these ligands and confers endocrine function. Klotho/betaKlotho have evolved as a compensatory mechanism for the poor ability of heparin/heparan sulfate to promote binding of FGF19, -21, and -23 to their cognate receptors.
- Department of Pharmacology, New York University School of Medicine, New York, NY 10016, USA.
Organizational Affiliation: