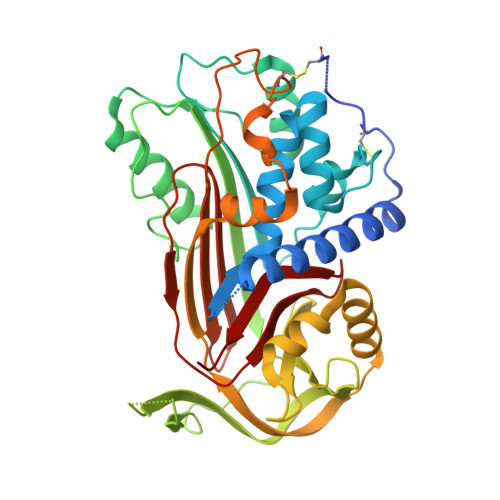
C1 inhibitor serpin domain structure reveals the likely mechanism of heparin potentiation and conformational disease
Beinrohr, L., Harmat, V., Dobo, J., Lorincz, Z., Gal, P., Zavodszky, P.(2007) J Biological Chem 282: 21100-21109
- PubMed: 17488724 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M700841200
- Primary Citation Related Structures:
2OAY - PubMed Abstract:
C1 inhibitor, a member of the serpin family, is a major down-regulator of inflammatory processes in blood. Genetic deficiency of C1 inhibitor results in hereditary angioedema, a dominantly inheritable, potentially lethal disease. Here we report the first crystal structure of the serpin domain of human C1 inhibitor, representing a previously unreported latent form, which explains functional consequences of several naturally occurring mutations, two of which are discussed in detail. The presented structure displays a novel conformation with a seven-stranded beta-sheet A. The unique conformation of the C-terminal six residues suggests its potential role as a barrier in the active-latent transition. On the basis of surface charge pattern, heparin affinity measurements, and docking of a heparin disaccharide, a heparin binding site is proposed in the contact area of the serpin-proteinase encounter complex. We show how polyanions change the activity of the C1 inhibitor by a novel "sandwich" mechanism, explaining earlier reaction kinetic and mutagenesis studies. These results may help to improve therapeutic C1 inhibitor preparations used in the treatment of hereditary angioedema, organ transplant rejection, and heart attack.
- Institute of Enzymology, Biological Research Center, Hungarian Academy of Sciences, Karolina út 29, H-1113 Budapest, Hungary. lbeinrohr@enzim.hu
Organizational Affiliation: