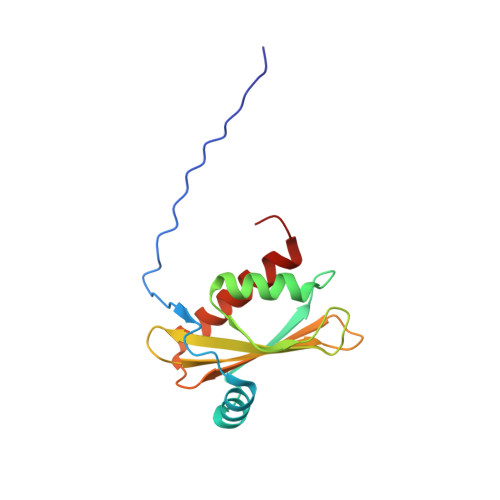
Solution NMR structure of beta-adaptin appendage domain of human adaptor protein complex 4 subunit beta, Northeast Structural Genomics Consortium (NESG) Target HR8998C
Eletsky, A., Rotshteyn, D.J., Pederson, K., Shastry, R., Maglaqui, M., Janjua, H., Xiao, R., Everett, J.K., Montelione, G.T., Prestegard, J.H., Szyperski, T.To be published.