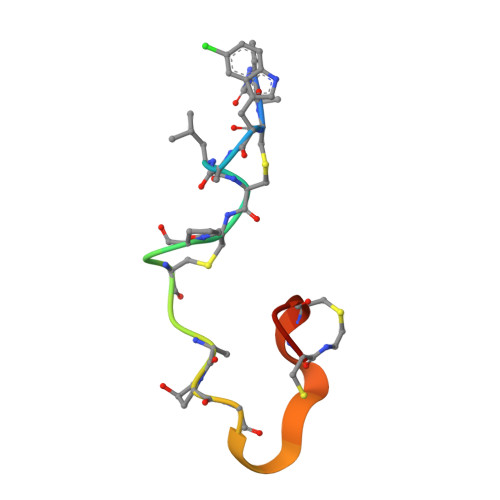
The Lantibiotic NAI-107 Binds to Bactoprenol-bound Cell Wall Precursors and Impairs Membrane Functions.
Munch, D., Muller, A., Schneider, T., Kohl, B., Wenzel, M., Bandow, J.E., Maffioli, S., Sosio, M., Donadio, S., Wimmer, R., Sahl, H.G.(2014) J Biological Chem 289: 12063-12076
- PubMed: 24627484
- DOI: https://doi.org/10.1074/jbc.M113.537449
- Primary Citation of Related Structures:
2MH5 - PubMed Abstract:
The lantibiotic NAI-107 is active against Gram-positive bacteria including vancomycin-resistant enterococci and methicillin-resistant Staphylococcus aureus. To identify the molecular basis of its potency, we studied the mode of action in a series of whole cell and in vitro assays and analyzed structural features by nuclear magnetic resonance (NMR). The lantibiotic efficiently interfered with late stages of cell wall biosynthesis and induced accumulation of the soluble peptidoglycan precursor UDP-N-acetylmuramic acid-pentapeptide (UDP-MurNAc-pentapeptide) in the cytoplasm. Using membrane preparations and a complete cascade of purified, recombinant late stage peptidoglycan biosynthetic enzymes (MraY, MurG, FemX, PBP2) and their respective purified substrates, we showed that NAI-107 forms complexes with bactoprenol-pyrophosphate-coupled precursors of the bacterial cell wall. Titration experiments indicate that first a 1:1 stoichiometric complex occurs, which then transforms into a 2:1 (peptide: lipid II) complex, when excess peptide is added. Furthermore, lipid II and related molecules obviously could not serve as anchor molecules for the formation of defined and stable nisin-like pores, however, slow membrane depolarization was observed after NAI-107 treatment, which could contribute to killing of the bacterial cell.
- Institute of Medical Microbiology, Immunology and Parasitology, Pharmaceutical Microbiology Section, University of Bonn, 53115 Bonn, Germany. Electronic address: dmuench@uni-bonn.de.
Organizational Affiliation: