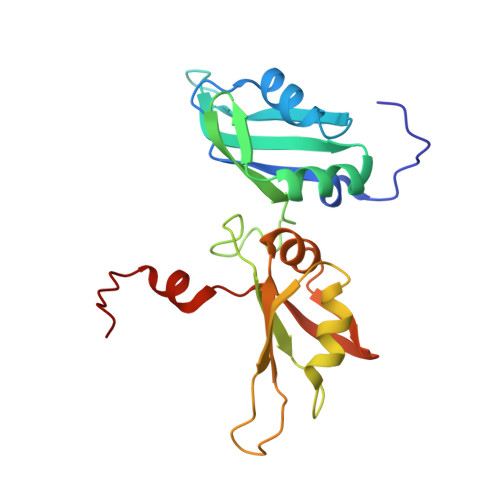
Solution structure of the two RNA recognition motifs of hnRNP A1 using segmental isotope labeling: how the relative orientation between RRMs influences the nucleic acid binding topology.
Barraud, P., Allain, F.H.(2013) J Biomol NMR 55: 119-138
- PubMed: 23247503 Search on PubMed
- DOI: https://doi.org/10.1007/s10858-012-9696-4
- Primary Citation Related Structures:
2LYV - PubMed Abstract:
Human hnRNP A1 is a multi-functional protein involved in many aspects of nucleic-acid processing such as alternative splicing, micro-RNA biogenesis, nucleo-cytoplasmic mRNA transport and telomere biogenesis and maintenance. The N-terminal region of hnRNP A1, also named unwinding protein 1 (UP1), is composed of two closely related RNA recognition motifs (RRM), and is followed by a C-terminal glycine rich region. Although crystal structures of UP1 revealed inter-domain interactions between RRM1 and RRM2 in both the free and bound form of UP1, these interactions have never been established in solution. Moreover, the relative orientation of hnRNP A1 RRMs is different in the free and bound crystal structures of UP1, raising the question of the biological significance of this domain movement. In the present study, we have used NMR spectroscopy in combination with segmental isotope labeling techniques to carefully analyze the inter-RRM contacts present in solution and subsequently determine the structure of UP1 in solution. Our data unambiguously demonstrate that hnRNP A1 RRMs interact in solution, and surprisingly, the relative orientation of the two RRMs observed in solution is different from the one found in the crystal structure of free UP1 and rather resembles the one observed in the nucleic-acid bound form of the protein. This strongly supports the idea that the two RRMs of hnRNP A1 have a single defined relative orientation which is the conformation previously observed in the bound form and now observed in solution using NMR. It is likely that the conformation in the crystal structure of the free form is a less stable form induced by crystal contacts. Importantly, the relative orientation of the RRMs in proteins containing multiple-RRMs strongly influences the RNA binding topologies that are practically accessible to these proteins. Indeed, RRM domains are asymmetric binding platforms contacting single-stranded nucleic acids in a single defined orientation. Therefore, the path of the nucleic acid molecule on the multiple RRM domains is strongly dependent on whether the RRMs are interacting with each other. The different nucleic acid recognition modes by multiple-RRM domains are briefly reviewed and analyzed on the basis of the current structural information.
- Institute of Molecular Biology and Biophysics, ETH Zurich, Schafmattstrasse 20, 8093 Zurich, Switzerland.
Organizational Affiliation: