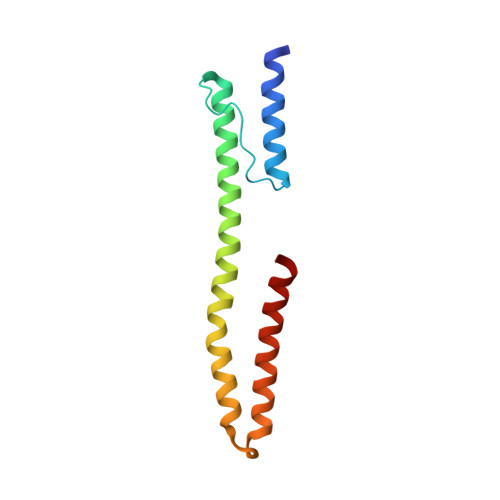
Mechanism of regulation of receptor histidine kinases.
Ferris, H.U., Dunin-Horkawicz, S., Hornig, N., Hulko, M., Martin, J., Schultz, J.E., Zeth, K., Lupas, A.N., Coles, M.(2012) Structure 20: 56-66
- PubMed: 22244755 Search on PubMed
- DOI: https://doi.org/10.1016/j.str.2011.11.014
- Primary Citation Related Structures:
2LFR, 2LFS, 3ZRV, 3ZRW, 3ZRX - PubMed Abstract:
Bacterial transmembrane receptors regulate an intracellular catalytic output in response to extracellular sensory input. To investigate the conformational changes that relay the regulatory signal, we have studied the HAMP domain, a ubiquitous intracellular module connecting input to output domains. HAMP forms a parallel, dimeric, four-helical coiled coil, and rational substitutions in our model domain (Af1503 HAMP) induce a transition in its interhelical packing, characterized by axial rotation of all four helices (the gearbox signaling model). We now illustrate how these conformational changes are propagated to a downstream domain by fusing Af1503 HAMP variants to the DHp domain of EnvZ, a bacterial histidine kinase. Structures of wild-type and mutant constructs are correlated with ligand response in vivo, clearly associating them with distinct signaling states. We propose that altered recognition of the catalytic domain by DHp, rather than a shift in position of the phospho-accepting histidine, forms the basis for regulation of kinase activity.
- Department of Protein Evolution, Max-Planck-Institute for Developmental Biology, 72076 Tübingen, Germany.
Organizational Affiliation: