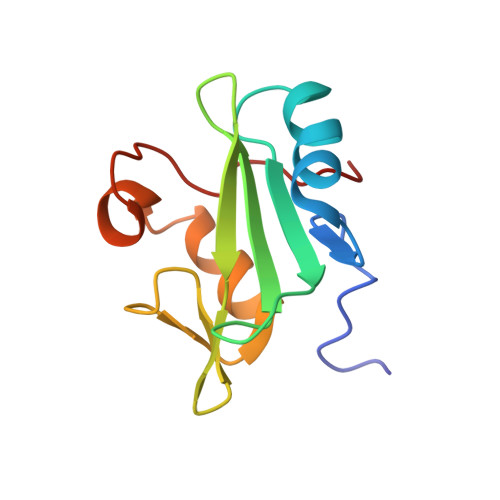
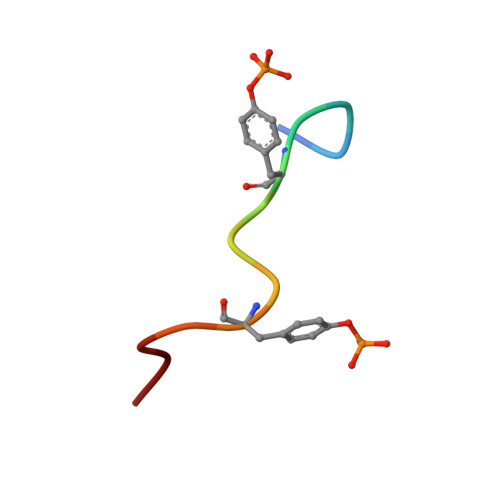
Two closely spaced tyrosines regulate NFAT signaling in B cells via Syk association with Vav.
Chen, C.H., Martin, V.A., Gorenstein, N.M., Geahlen, R.L., Post, C.B.(2011) Mol Cell Biol 31: 2984-2996
- PubMed: 21606197 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1128/MCB.05043-11
- Primary Citation Related Structures:
2LCT - PubMed Abstract:
Activated Syk, an essential tyrosine kinase in B cell signaling, interacts with Vav guanine nucleotide exchange factors and regulates Vav activity through tyrosine phosphorylation. The Vav SH2 domain binds Syk linker B by an unusual recognition of two closely spaced Syk tyrosines: Y342 and Y346. The binding affinity is highest when both Y342 and Y346 are phosphorylated. An investigation in B cells of the dependence of Vav phosphorylation and NFAT activation on phosphorylation of Y342 and Y346 finds that cellular response levels match the relative binding affinities of the Vav1 SH2 domain for singly and doubly phosphorylated linker B peptides. This key result suggests that the uncommon recognition determinant of these two closely spaced tyrosines is a limiting factor in signaling. Interestingly, differences in affinities for binding singly and doubly phosphorylated peptides are reflected in the on rate, not the off rate. Such a control mechanism would be highly effective for regulating binding among competing Syk binding partners. The nuclear magnetic resonance (NMR) structure of Vav1 SH2 in complex with a doubly phosphorylated linker B peptide reveals diverse conformations associated with the unusual SH2 recognition of two phosphotyrosines. NMR relaxation indicates compensatory changes in loop fluctuations upon binding, with implications for nonphosphotyrosine interactions of Vav1 SH2.
- Department of Medicinal Chemistry and Molecular Pharmacology, Markey Center for Structural Biology and Purdue Center for Cancer Research, Purdue University, West Lafayette, Indiana 47907, USA.
Organizational Affiliation: