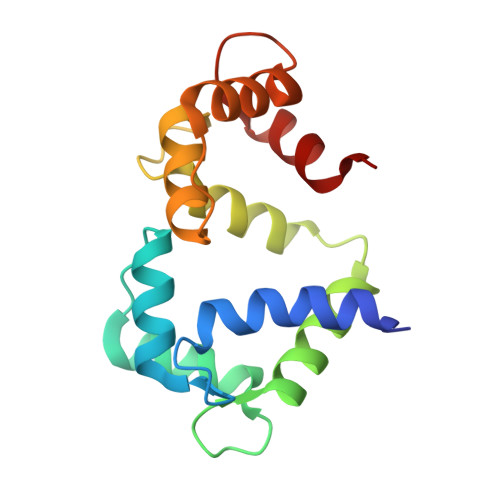
Structure of Calmodulin Bound to the Hydrophobic IQ Domain of the Cardiac Ca(v)1.2 Calcium Channel.
Fallon, J.L., Halling, D.B., Hamilton, S.L., Quiocho, F.A.(2005) Structure 13: 1881-1886
- PubMed: 16338416 Search on PubMed
- DOI: https://doi.org/10.1016/j.str.2005.09.021
- Primary Citation Related Structures:
2F3Y, 2F3Z - PubMed Abstract:
Ca2+-dependent inactivation (CDI) and facilitation (CDF) of the Ca(v)1.2 Ca2+ channel require calmodulin binding to a putative IQ motif in the carboxy-terminal tail of the pore-forming subunit. We present the 1.45 A crystal structure of Ca2+-calmodulin bound to a 21 residue peptide corresponding to the IQ domain of Ca(v)1.2. This structure shows that parallel binding of calmodulin to the IQ domain is governed by hydrophobic interactions. Mutations of residues I1672 and Q1673 in the peptide to alanines, which abolish CDI but not CDF in the channel, do not greatly alter the structure. Both lobes of Ca2+-saturated CaM bind to the IQ peptide but isoleucine 1672, thought to form an intramolecular interaction that drives CDI, is buried. These findings suggest that this structure could represent the conformation that calmodulin assumes in CDF.
- Verna and Marrs McLean Department of Biochemistry and Molecular Biology, Baylor College of Medicine, Houston, Texas 77030, USA.
Organizational Affiliation: