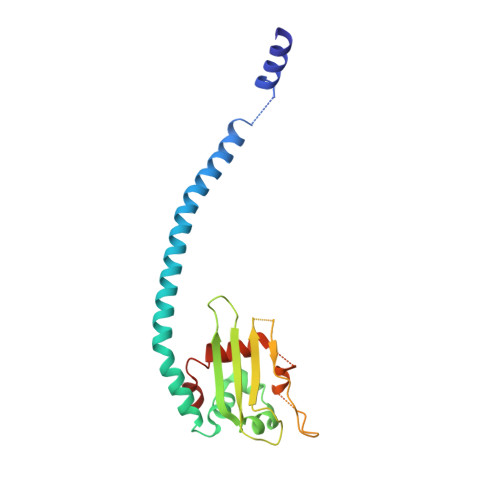
Relationship between the structure of SET/TAF-Ibeta/INHAT and its histone chaperone activity
Muto, S., Senda, M., Akai, Y., Sato, L., Suzuki, T., Nagai, R., Senda, T., Horikoshi, M.(2007) Proc Natl Acad Sci U S A 104: 4285-4290
- PubMed: 17360516 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.0603762104
- Primary Citation Related Structures:
2E50 - PubMed Abstract:
Histone chaperones assemble and disassemble nucleosomes in an ATP-independent manner and thus regulate the most fundamental step in the alteration of chromatin structure. The molecular mechanisms underlying histone chaperone activity remain unclear. To gain insights into these mechanisms, we solved the crystal structure of the functional domain of SET/TAF-Ibeta/INHAT at a resolution of 2.3 A. We found that SET/TAF-Ibeta/INHAT formed a dimer that assumed a "headphone"-like structure. Each subunit of the SET/TAF-Ibeta/INHAT dimer consisted of an N terminus, a backbone helix, and an "earmuff" domain. It resembles the structure of the related protein NAP-1. Comparison of the crystal structures of SET/TAF-Ibeta/INHAT and NAP-1 revealed that the two proteins were folded similarly except for an inserted helix. However, their backbone helices were shaped differently, and the relative dispositions of the backbone helix and the earmuff domain between the two proteins differed by approximately 40 degrees . Our biochemical analyses of mutants revealed that the region of SET/TAF-Ibeta/INHAT that is engaged in histone chaperone activity is the bottom surface of the earmuff domain, because this surface bound both core histones and double-stranded DNA. This overlap or closeness of the activity surface and the binding surfaces suggests that the specific association among SET/TAF-Ibeta/INHAT, core histones, and double-stranded DNA is requisite for histone chaperone activity. These findings provide insights into the possible mechanisms by which histone chaperones assemble and disassemble nucleosome structures.
- Laboratory of Developmental Biology, Institute of Molecular and Cellular Biosciences, University of Tokyo, 1-1-1 Yayoi, Bunkyo-ku, Tokyo 113-0032, Japan.
Organizational Affiliation: