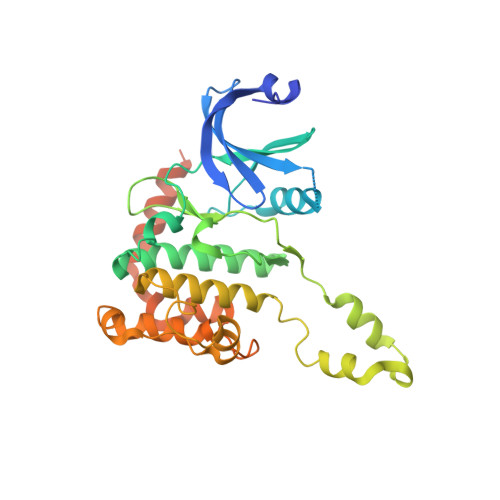
Trans-Activation of the DNA-Damage Signalling Protein Kinase Chk2 by T-Loop Exchange
Oliver, A.W., Paul, A., Boxall, K.J., Barrie, S.E., Aherne, G.W., Garrett, M.D., Mittnacht, S., Pearl, L.H.(2006) EMBO J 25: 3179
- PubMed: 16794575 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/sj.emboj.7601209
- Primary Citation Related Structures:
2CN5, 2CN8 - PubMed Abstract:
The protein kinase Chk2 (checkpoint kinase 2) is a major effector of the replication checkpoint. Chk2 activation is initiated by phosphorylation of Thr68, in the serine-glutamine/threonine-glutamine cluster domain (SCD), by ATM. The phosphorylated SCD-segment binds to the FHA domain of a second Chk2 molecule, promoting dimerisation of the protein and triggering phosphorylation of the activation segment/T-loop in the kinase domain. We have now determined the structure of the kinase domain of human Chk2 in complexes with ADP and a small-molecule inhibitor debromohymenialdisine. The structure reveals a remarkable dimeric arrangement in which T-loops are exchanged between protomers, to form an active kinase conformation in trans. Biochemical data suggest that this dimer is the biologically active state promoted by ATM-phosphorylation, and also suggests a mechanism for dimerisation-driven activation of Chk2 by trans-phosphorylation.
- Section of Structural Biology, Cancer Research UK DNA Repair Enzymes Group, The Institute of Cancer Research, Chelsea, London, UK. antony.oliver@icr.ac.uk
Organizational Affiliation: