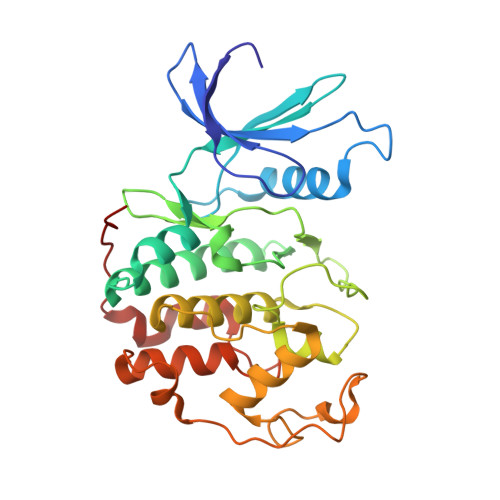
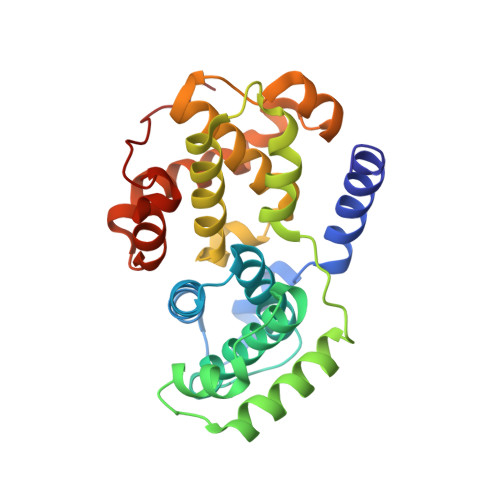
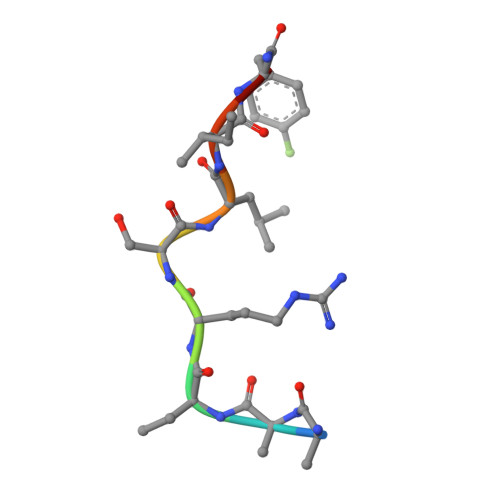
Differential Binding of Inhibitors to Active and Inactive Cdk2 Provides Insights for Drug Design.
Kontopidis, G., Mcinnes, C., Pandalaneni, S.R., Mcnae, I., Gibson, D., Mezna, M., Thomas, M., Wood, G., Wang, S., Walkinshaw, M.D., Fischer, P.M.(2006) Chem Biol 13: 201
- PubMed: 16492568 Search on PubMed
- DOI: https://doi.org/10.1016/j.chembiol.2005.11.011
- Primary Citation Related Structures:
2C5N, 2C5O, 2C5V, 2C5X, 2C5Y - PubMed Abstract:
The cyclin-dependent kinases (CDKs) have been characterized in complex with a variety of inhibitors, but the majority of structures solved are in the inactive form. We have solved the structures of six inhibitors in both the monomeric CDK2 and binary CDK2/cyclinA complexes and demonstrate that significant differences in ligand binding occur depending on the activation state. The binding mode of two ligands in particular varies substantially in active and inactive CDK2. Furthermore, energetic analysis of CDK2/cyclin/inhibitors demonstrates that a good correlation exists between the in vitro potency and the calculated energies of interaction, but no such relationship exists for CDK2/inhibitor structures. These results confirm that monomeric CDK2 ligand complexes do not fully reflect active conformations, revealing significant implications for inhibitor design while also suggesting that the monomeric CDK2 conformation can be selectively inhibited.
- Cyclacel Ltd., Dundee, UK.
Organizational Affiliation: