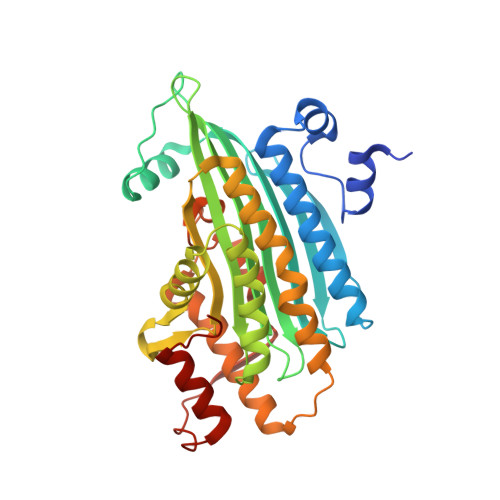
Structural basis of hereditary coproporphyria.
Lee, D.S., Flachsova, E., Bodnarova, M., Demeler, B., Martasek, P., Raman, C.S.(2005) Proc Natl Acad Sci U S A 102: 14232-14237
- PubMed: 16176984 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.0506557102
- Primary Citation Related Structures:
2AEX - PubMed Abstract:
Hereditary coproporphyria is an autosomal dominant disorder resulting from the half-normal activity of coproporphyrinogen oxidase (CPO), a mitochondrial enzyme catalyzing the antepenultimate step in heme biosynthesis. The mechanism by which CPO catalyzes oxidative decarboxylation, in an extraordinary metal- and cofactor-independent manner, is poorly understood. Here, we report the crystal structure of human CPO at 1.58-A resolution. The structure reveals a previously uncharacterized tertiary topology comprising an unusually flat seven-stranded beta-sheet sandwiched by alpha-helices. In the biologically active dimer (K(D) = 5 x 10(-7) M), one monomer rotates relative to the second by approximately 40 degrees to create an intersubunit interface in close proximity to two independent enzymatic sites. The unexpected finding of citrate at the active site allows us to assign Ser-244, His-258, Asn-260, Arg-262, Asp-282, and Arg-332 as residues mediating substrate recognition and decarboxylation. We favor a mechanism in which oxygen serves as the immediate electron acceptor, and a substrate radical or a carbanion with substantial radical character participates in catalysis. Although several mutations in the CPO gene have been described, the molecular basis for how these alterations diminish enzyme activity is unknown. We show that deletion of residues (392-418) encoded by exon six disrupts dimerization. Conversely, harderoporphyria-causing K404E mutation precludes a type I beta-turn from retaining the substrate for the second decarboxylation cycle. Together, these findings resolve several questions regarding CPO catalysis and provide insights into hereditary coproporphyria.
- Department of Biochemistry and Molecular Biology, University of Texas Medical School, Houston, TX 77030, USA.
Organizational Affiliation: