Antiproliferative activity of olomoucine II, a novel 2,6,9-trisubstituted purine cyclin-dependent kinase inhibitor
Krystof, V., McNae, I.W., Walkinshaw, M.D., Fischer, P.M., Muller, P., Vojtesek, B., Orsag, M., Havlicek, L., Strnad, M.(2005) Cell Mol Life Sci 62: 1763-1771
- PubMed: 16003486 Search on PubMed
- DOI: https://doi.org/10.1007/s00018-005-5185-1
- Primary Citation Related Structures:
2A0C - PubMed Abstract:
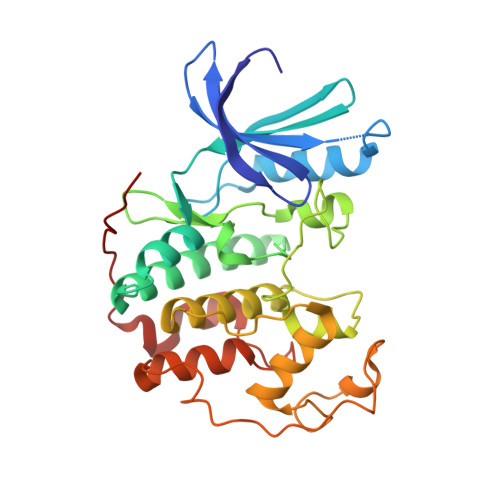
The study describes the protein kinase selectivity profile, as well as the binding mode of olomoucine II in the catalytic cleft of CDK2, as determined from cocrystal analysis. Apart from the main cell cycle-regulating kinase CDK2, olomoucine II exerts specificity for CDK7 and CDK9, with important functions in the regulation of RNA transcription. In vitro anticancer activity of the inhibitor in a panel of tumor cell lines shows a wide potency range with a slight preference for cells harboring a wild-type p53 gene. Cell-based assays confirmed activation of p53 protein levels and events leading to accumulation of p21(WAF1). Additionally, in olomoucine II-treated cells, Mdm2 was found to form a complex with the ribosomal protein L11, which inhibits Mdm2 ubiquitin ligase function. We conclude that perturbations in RNA synthesis may lead to activation of p53 and that this contributes to the antiproliferative potency of cyclindependent kinase inhibitors.
- Laboratory of Growth Regulators, Faculty of Science, Palacký University & Institute of Experimental Botany AS CR, Slechtitelů 11, 783 71, Olomouc, The Czech Republic.
Organizational Affiliation: