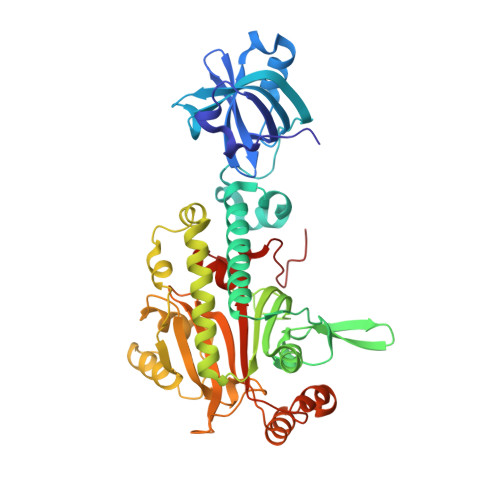
Structural Basis of the Water-assisted Asparagine Recognition by Asparaginyl-tRNA Synthetase.
Iwasaki, W., Sekine, S., Kuroishi, C., Kuramitsu, S., Shirouzu, M., Yokoyama, S.(2006) J Mol Biology 360: 329-342
- PubMed: 16753178 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2006.04.068
- Primary Citation Related Structures:
1X54, 1X55, 1X56 - PubMed Abstract:
Asparaginyl-tRNA synthetase (AsnRS) is a member of the class-II aminoacyl-tRNA synthetases, and is responsible for catalyzing the specific aminoacylation of tRNA(Asn) with asparagine. Here, the crystal structure of AsnRS from Pyrococcus horikoshii, complexed with asparaginyl-adenylate (Asn-AMP), was determined at 1.45 A resolution, and those of free AsnRS and AsnRS complexed with an Asn-AMP analog (Asn-SA) were solved at 1.98 and 1.80 A resolutions, respectively. All of the crystal structures have many solvent molecules, which form a network of hydrogen-bonding interactions that surrounds the entire AsnRS molecule. In the AsnRS/Asn-AMP complex (or the AsnRS/Asn-SA), one side of the bound Asn-AMP (or Asn-SA) is completely covered by the solvent molecules, which complement the binding site. In particular, two of these water molecules were found to interact directly with the asparagine amide and carbonyl groups, respectively, and to contribute to the formation of a pocket highly complementary to the asparagine side-chain. Thus, these two water molecules appear to play a key role in the strict recognition of asparagine and the discrimination against aspartic acid by the AsnRS. This water-assisted asparagine recognition by the AsnRS strikingly contrasts with the fact that the aspartic acid recognition by the closely related aspartyl-tRNA synthetase is achieved exclusively through extensive interactions with protein amino acid residues. Furthermore, based on a docking model of AsnRS and tRNA, a single arginine residue (Arg83) in the AsnRS was postulated to be involved in the recognition of the third position of the tRNA(Asn) anticodon (U36). We performed a mutational analysis of this particular arginine residue, and confirmed its significance in the tRNA recognition.
- Department of Biophysics and Biochemistry, Graduate School of Science, The University of Tokyo, 7-3-1 Hongo, Bunkyo-ku, Tokyo 113-0033, Japan.
Organizational Affiliation: