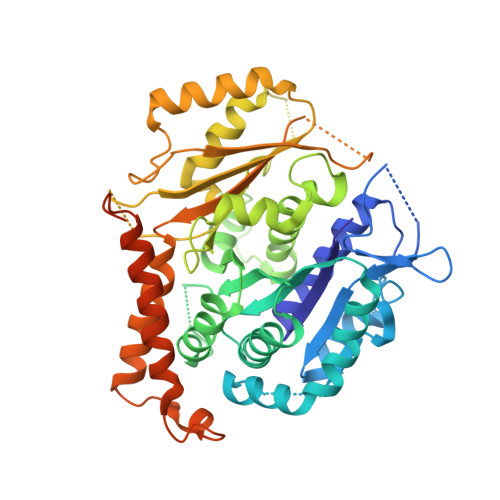
Insights into microtubule nucleation from the crystal structure of human gamma-tubulin.
Aldaz, H., Rice, L.M., Stearns, T., Agard, D.A.(2005) Nature 435: 523-527
- PubMed: 15917813 Search on PubMed
- DOI: https://doi.org/10.1038/nature03586
- Primary Citation Related Structures:
1Z5V, 1Z5W - PubMed Abstract:
Microtubules are hollow polymers of alphabeta-tubulin that show GTP-dependent assembly dynamics and comprise a critical part of the eukaryotic cytoskeleton. Initiation of new microtubules in vivo requires gamma-tubulin, organized as an oligomer within the 2.2-MDa gamma-tubulin ring complex (gamma-TuRC) of higher eukaryotes. Structural insight is lacking regarding gamma-tubulin, its oligomerization and how it promotes microtubule assembly. Here we report the 2.7-A crystal structure of human gamma-tubulin bound to GTP-gammaS (a non-hydrolysable GTP analogue). We observe a 'curved' conformation for gamma-tubulin-GTPgammaS, similar to that seen for GDP-bound, unpolymerized alphabeta-tubulin. Tubulins are thought to represent a distinct class of GTP-binding proteins, and conformational switching in gamma-tubulin might differ from the nucleotide-dependent switching of signalling GTPases. A crystal packing interaction replicates the lateral contacts between alpha- and beta-tubulins in the microtubule, and this association probably forms the basis for gamma-tubulin oligomerization within the gamma-TuRC. Laterally associated gamma-tubulins in the gamma-TuRC might promote microtubule nucleation by providing a template that enhances the intrinsically weak lateral interaction between alphabeta-tubulin heterodimers. Because they are dimeric, alphabeta-tubulins cannot form microtubule-like lateral associations in the curved conformation. The lateral array of gamma-tubulins we observe in the crystal reveals a unique functional property of a monomeric tubulin.
- Howard Hughes Medical Institute and Department of Biochemistry and Biophysics, University of California, San Francisco, 600 16th Street, San Francisco, California 94143, USA.
Organizational Affiliation: