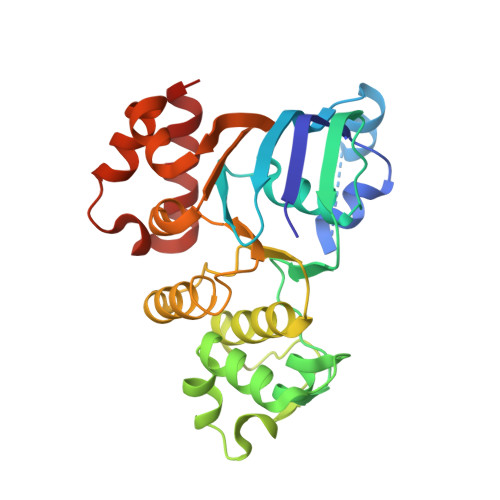
Side chain and backbone contributions of Phe508 to CFTR folding.
Thibodeau, P.H., Brautigam, C.A., Machius, M., Thomas, P.J.(2005) Nat Struct Mol Biol 12: 10-16
- PubMed: 15619636 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/nsmb881
- Primary Citation Related Structures:
1XF9, 1XFA - PubMed Abstract:
Mutations in the cystic fibrosis transmembrane conductance regulator (CFTR), an integral membrane protein, cause cystic fibrosis (CF). The most common CF-causing mutant, deletion of Phe508, fails to properly fold. To elucidate the role Phe508 plays in the folding of CFTR, missense mutations at this position were generated. Only one missense mutation had a pronounced effect on the stability and folding of the isolated domain in vitro. In contrast, many substitutions, including those of charged and bulky residues, disrupted folding of full-length CFTR in cells. Structures of two mutant nucleotide-binding domains (NBDs) reveal only local alterations of the surface near position 508. These results suggest that the peptide backbone plays a role in the proper folding of the domain, whereas the side chain plays a role in defining a surface of NBD1 that potentially interacts with other domains during the maturation of intact CFTR.
- Department of Physiology, The University of Texas Southwestern Medical Center at Dallas, 75390 USA.
Organizational Affiliation: