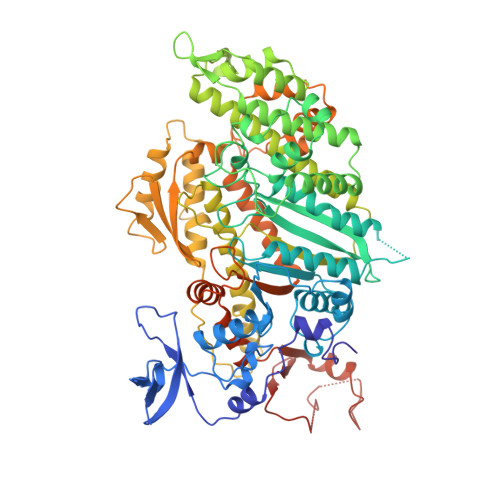
X-ray structure of the magnesium(II).ADP.vanadate complex of the Dictyostelium discoideum myosin motor domain to 1.9 A resolution.
Smith, C.A., Rayment, I.(1996) Biochemistry 35: 5404-5417
- PubMed: 8611530 Search on PubMed
- DOI: https://doi.org/10.1021/bi952633+
- Primary Citation Related Structures:
1VOM - PubMed Abstract:
The structure of the vanadate-trapped ADP complex of a truncated head of Dictyostelium myosin II consisting of residues Asp 2-Asn 762 has been determined by molecular replacement at 1.9 A resolution and refined to a crystallographic R-factor of 19.4%. The crystals belong to the orthorhombic space group C2221 where a = 84.50 A, b = 145.4 A, and c = 152.8 A. The conformation of the protein is similar to that of MgADP.AlF4.SlDc [Fisher, A.J., et al. (1995) Biochemistry 34, 8960-8972]. The nucleotide binding site contains a complex between MgADP and vanadate where MgADP exhibits a very similar conformation to that seen in previous complexes. The vanadate ion adopts a trigonal bipyramidal coordination. The three equatorial oxygen ligands are fairly short, average 1.7 A, relative to a single bond distance of approximately 1.8 A and are coordinated to the magnesium ion, N zeta of Lys 185, and five other protein ligands. The apical coordination to the vanadate ion is filled by a terminal oxygen on the beta-phosphate of ADP and a water molecule at bond distances of 2.1 and 2.3 A, respectively. The long length of the apical bonds suggests that the bond order is considerably less than unity. This structure confirms the earlier suggestion that vanadate is a model for the transition state of ATP hydrolysis and thus provides insight into those factors that are responsible for catalysis. In particular, it shows that the protein ligands and water structure surrounding the gamma-phosphate pocket are oriented to stabilize a water molecule in an appropriate position for in-line nucleophilic attack on the gamma-phosphorus of ATP. This structure reveals also an orientation of the COOH-terminal region beyond Thr 688 which is very different from that observed in either MgADP.BeFx.SlDc or chicken skeletal myosin subfragment 1. This is consistent with the COOH-terminal region of the molecule playing an important role in the transduction of chemical energy of hydrolysis of ATP into mechanical movement.
- Institute for Enzyme Research, University of Wisconsin, Madison 53705, USA.
Organizational Affiliation: