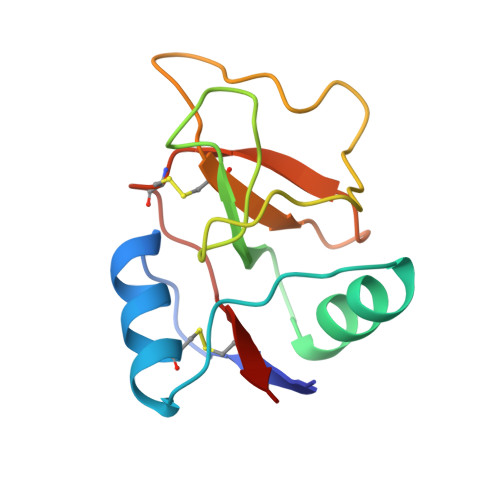
Structural analysis of monosaccharide recognition by rat liver mannose-binding protein.
Ng, K.K., Drickamer, K., Weis, W.I.(1996) J Biological Chem 271: 663-674
- PubMed: 8557671 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.271.2.663
- Primary Citation Related Structures:
1RDI, 1RDJ, 1RDK, 1RDL, 1RDM, 1RDN, 1RDO - PubMed Abstract:
The structural basis of carbohydrate recognition by rat liver mannose-binding protein (MBP-C) has been explored by determining the three-dimensional structure of the C-type carbohydrate-recognition domain (CRD) of MBP-C using x-ray crystallography. The structure was solved by molecular replacement using rat serum mannose-binding protein (MBP-A) as a search model and was refined to maximum Bragg spacings of 1.7 A. Despite their almost identical folds, the dimeric structures formed by the two MBP CRDs differ dramatically. Complexes of MBP-C with methyl glycosides of mannose, N-acetylglucosamine, and fucose were prepared by soaking MBP-C crystals in solutions containing these sugars. Surprisingly, the pyranose ring of mannose is rotated 180 degrees relative to the orientation observed previously in MBP-A, but the local interactions between sugar and protein are preserved. For each of the bound sugars, vicinal, equatorial hydroxyl groups equivalent to the 3- and 4-OH groups of mannose directly coordinate Ca2+ and form hydrogen bonds with residues also serving as Ca2+ ligands. Few interactions are observed between other parts of the sugar and the protein. A complex formed between free galactose and MBP-C reveals a similar mode of binding, with the anomeric hydroxyl group serving as one of the Ca2+ ligands. A second binding site for mannose has also been observed in one of two copies in the asymmetric unit at a sugar concentration of 1.3 M. These structures explain how MBPs recognize a wide range of monosaccharides and suggest how fine specificity differences between MBP-A and MBP-C may be achieved.
- Department of Structural Biology, Stanford University School of Medicine, California 94305, USA.
Organizational Affiliation: