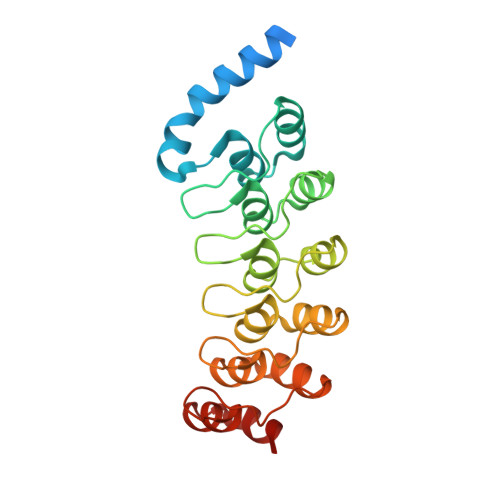
Structure and stability of the ankyrin domain of the Drosophila Notch receptor
Zweifel, M.E., Leahy, D.J., Hughson, F.M., Barrick, D.(2003) Protein Sci 12: 2622-2632
- PubMed: 14573873 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1110/ps.03279003
- Primary Citation Related Structures:
1OT8 - PubMed Abstract:
The Notch receptor contains a conserved ankyrin repeat domain that is required for Notch-mediated signal transduction. The ankyrin domain of Drosophila Notch contains six ankyrin sequence repeats previously identified as closely matching the ankyrin repeat consensus sequence, and a putative seventh C-terminal sequence repeat that exhibits lower similarity to the consensus sequence. To better understand the role of the Notch ankyrin domain in Notch-mediated signaling and to examine how structure is distributed among the seven ankyrin sequence repeats, we have determined the crystal structure of this domain to 2.0 angstroms resolution. The seventh, C-terminal, ankyrin sequence repeat adopts a regular ankyrin fold, but the first, N-terminal ankyrin repeat, which contains a 15-residue insertion, appears to be largely disordered. The structure reveals a substantial interface between ankyrin polypeptides, showing a high degree of shape and charge complementarity, which may be related to homotypic interactions suggested from indirect studies. However, the Notch ankyrin domain remains largely monomeric in solution, demonstrating that this interface alone is not sufficient to promote tight association. Using the structure, we have classified reported mutations within the Notch ankyrin domain that are known to disrupt signaling into those that affect buried residues and those restricted to surface residues. We show that the buried substitutions greatly decrease protein stability, whereas the surface substitutions have only a marginal affect on stability. The surface substitutions are thus likely to interfere with Notch signaling by disrupting specific Notch-effector interactions and map the sites of these interactions.
- T.C. Jenkins Department of Biophysics, The Johns Hopkins University, Baltimore, Maryland 21218, USA.
Organizational Affiliation: