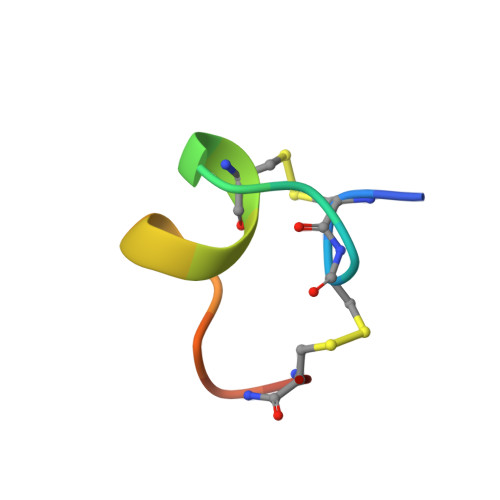
Three-dimensional solution structure of alpha-conotoxin MII by NMR spectroscopy: effects of solution environment on helicity.
Hill, J.M., Oomen, C.J., Miranda, L.P., Bingham, J.P., Alewood, P.F., Craik, D.J.(1998) Biochemistry 37: 15621-15630
- PubMed: 9843366 Search on PubMed
- DOI: https://doi.org/10.1021/bi981535w
- Primary Citation Related Structures:
1MII - PubMed Abstract:
alpha-Conotoxin MII, a 16-residue polypeptide from the venom of the piscivorous cone snail Conus magus, is a potent and highly specific blocker of mammalian neuronal nicotinic acetylcholine receptors composed of alpha3 beta2 subunits. The role of this receptor type in the modulation of neurotransmitter release and its relevance to the problems of addiction and psychosis emphasize the importance of a structural understanding of the mode of interaction of MII with the alpha3 beta2 interface. Here we describe the three-dimensional solution structure of MII determined using 2D 1H NMR spectroscopy. Structural restraints consisting of 376 interproton distances inferred from NOEs and 12 dihedral restraints derived from spin-spin coupling constants were used as input for simulated annealing calculations and energy minimization in the program X-PLOR. The final set of 20 structures is exceptionally well-defined with mean pairwise rms differences over the whole molecule of 0.07 A for the backbone atoms and 0.34 A for all heavy atoms. MII adopts a compact structure incorporating a central segment of alpha-helix and beta-turns at the N- and C-termini. The molecule is stabilized by two disulfide bonds, which provide cross-links between the N-terminus and both the middle and C-terminus of the structure. The susceptibility of the structure to conformational change was examined using several different solvent conditions. While the global fold of MII remains the same, the structure is stabilized in a more hydrophobic environment provided by the addition of acetonitrile or trifluoroethanol to the aqueous solution. The distribution of amino acid side chains in MII creates distinct hydrophobic and polar patches on its surface that may be important for the specific interaction with the alpha3beta2 neuronal nAChR. A comparison of the structure of MII with other neuronal-specific alpha-conotoxins provides insights into their mode of interaction with these receptors.
- Centre for Drug Design and Development, The University of Queensland, Brisbane, Australia.
Organizational Affiliation: