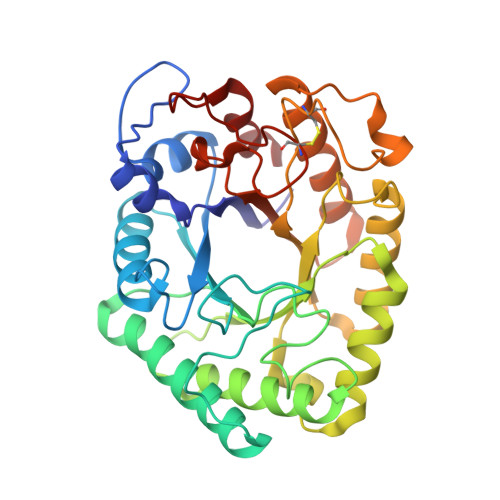
Structure of Two Fungal Beta-1,4-Galactanases: Searching for the Basis for Temperature and Ph Optimum
Le Nours, J., Ryttersgaard, C., Lo Leggio, L., Ostergaard, P.R., Borchert, T.V., Christensen, L.L.H., Larsen, S.(2003) Protein Sci 12: 1195
- PubMed: 12761390 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1110/ps.0300103
- Primary Citation Related Structures:
1HJQ, 1HJS, 1HJU - PubMed Abstract:
beta-1,4-Galactanases hydrolyze the galactan side chains that are part of the complex carbohydrate structure of the pectin. They are assigned to family 53 of the glycoside hydrolases and display significant variations in their pH and temperature optimum and stability. Two fungal beta-1,4-galactanases from Myceliophthora thermophila and Humicola insolens have been cloned and heterologously expressed, and the crystal structures of the gene products were determined. The structures are compared to the previously only known family 53 structure of the galactanase from Aspergillus aculeatus (AAGAL) showing approximately 56% identity. The M. thermophila and H. insolens galactanases are thermophilic enzymes and are most active at neutral to basic pH, whereas AAGAL is mesophilic and most active at acidic pH. The structure of the M. thermophila galactanase (MTGAL) was determined from crystals obtained with HEPES and TRIS buffers to 1.88 A and 2.14 A resolution, respectively. The structure of the H. insolens galactanase (HIGAL) was determined to 2.55 A resolution. The thermostability of MTGAL and HIGAL correlates with increase in the protein rigidity and electrostatic interactions, stabilization of the alpha-helices, and a tighter packing. An inspection of the active sites in the three enzymes identifies several amino acid substitutions that could explain the variation in pH optimum. Examination of the activity as a function of pH for the D182N mutant of AAGAL and the A90S/ H91D mutant of MTGAL showed that the difference in pH optimum between AAGAL and MTGAL is at least partially associated with differences in the nature of residues at positions 182, 90, and/or 91.
- Centre for Crystallographic Studies, Department of Chemistry, University of Copenhagen, DK-2100 Copenhagen, Denmark.
Organizational Affiliation: