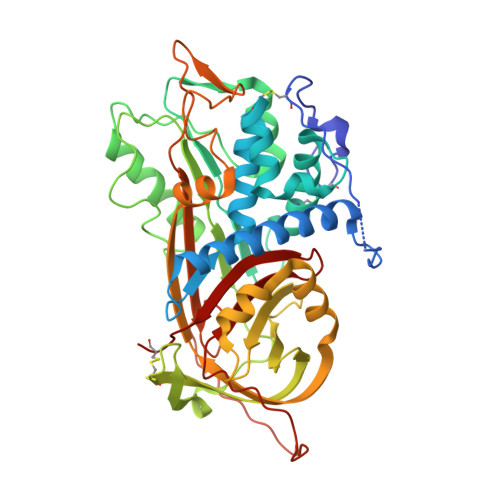
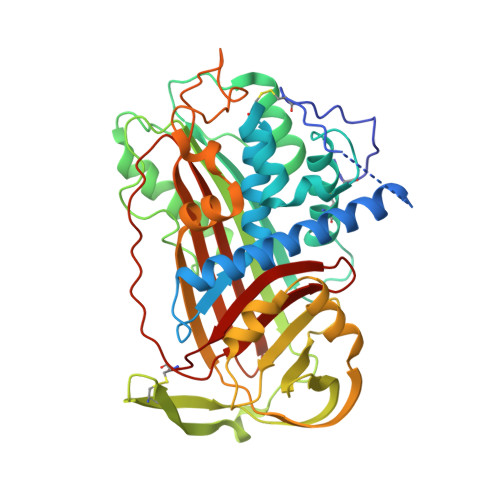
The Conformational Activation of Antithrombin. A 2. 85-A Structure of a Fluorescein Derivative Reveals an Electrostatic Link between the Hinge and Heparin Binding Regions.
Huntington, J.A., Mccoy, A.J., Belzar, K.J., Pei, X.Y., Gettins, P.G.W., Carrell, R.W.(2000) J Biological Chem 275: 15377
- PubMed: 10809774 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.275.20.15377
- Primary Citation Related Structures:
1DZG, 1DZH - PubMed Abstract:
Antithrombin is unique among the serpins in that it circulates in a native conformation that is kinetically inactive toward its target proteinase, factor Xa. Activation occurs upon binding of a specific pentasaccharide sequence found in heparin that results in a rearrangement of the reactive center loop removing constraints on the active center P1 residue. We determined the crystal structure of an activated antithrombin variant, N135Q S380C-fluorescein (P14-fluorescein), in order to see how full activation is achieved in the absence of heparin and how the structural effects of the substitution in the hinge region are translated to the heparin binding region. The crystal structure resembles native antithrombin except in the hinge and heparin binding regions. The absence of global conformational change allows for identification of specific interactions, centered on Glu(381) (P13), that are responsible for maintenance of the solution equilibrium between the native and activated forms and establishes the existence of an electrostatic link between the hinge region and the heparin binding region. A revised model for the mechanism of the allosteric activation of antithrombin is proposed.
- University of Cambridge, Department of Haematology, Wellcome Trust Centre for the Study of Molecular Mechanisms in Disease, Cambridge Institute for Medical Research, Hills Road, Cambridge CB2 2XY, United Kingdom. jah52@cam.ac.uk
Organizational Affiliation: