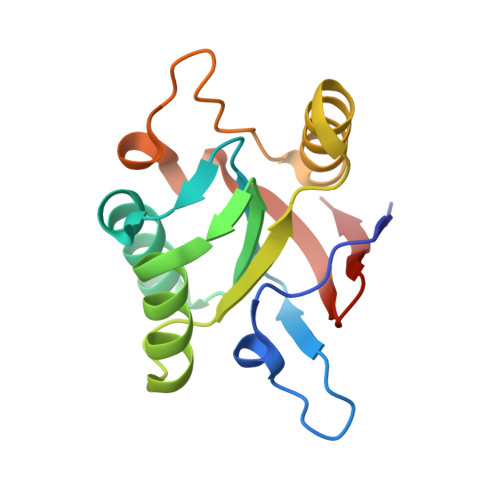
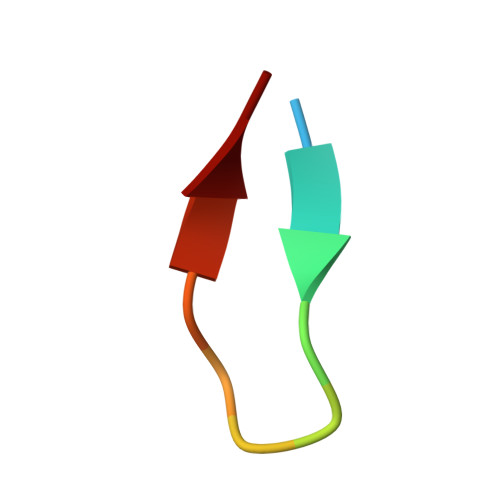
The crystal structure of a GroEL/peptide complex: plasticity as a basis for substrate diversity.
Chen, L., Sigler, P.B.(1999) Cell 99: 757-768
- PubMed: 10619429 Search on PubMed
- DOI: https://doi.org/10.1016/s0092-8674(00)81673-6
- Primary Citation Related Structures:
1DK7, 1DKD - PubMed Abstract:
The chaperonin GroEL is a double toriodal assembly that with its cochaperonin GroES facilitates protein folding with an ATP-dependent mechanism. Nonnative conformations of diverse protein substrates bind to the apical domains surrounding the opening of the double toroid's central cavity. Using phage display, we have selected peptides with high affinity for the isolated apical domain. We have determined the crystal structures of the complexes formed by the most strongly bound peptide with the isolated apical domain, and with GroEL. The peptide interacts with the groove between paired alpha helices in a manner similar to that of the GroES mobile loop. Our structural analysis, combined with other results, suggests that various modes of molecular plasticity are responsible for tight promiscuous binding of nonnative substrates and their release into the shielded cis assembly.
- Department of Molecular Biophysics and Biochemistry, Yale University, New Haven, Connecticut 06511, USA.
Organizational Affiliation: