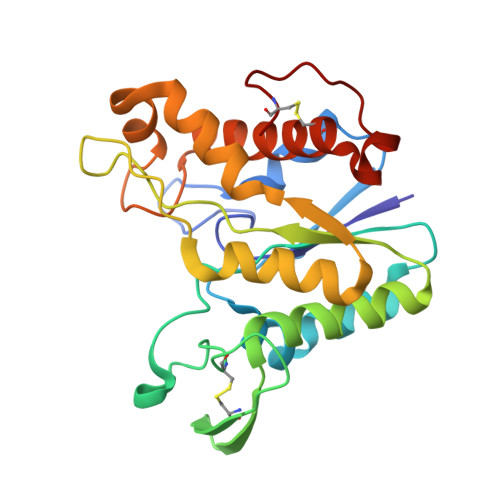
Rhamnogalacturonan acetylesterase elucidates the structure and function of a new family of hydrolases.
Molgaard, A., Kauppinen, S., Larsen, S.(2000) Structure 8: 373-383
- PubMed: 10801485 Search on PubMed
- DOI: https://doi.org/10.1016/s0969-2126(00)00118-0
- Primary Citation Related Structures:
1DEO, 1DEX - PubMed Abstract:
The complex polysaccharide rhamnogalacturonan constitutes a major part of the hairy region of pectin. It can have different types of carbohydrate sidechains attached to the rhamnose residues in the backbone of alternating rhamnose and galacturonic acid residues; the galacturonic acid residues can be methylated or acetylated. Aspergillus aculeatus produces enzymes that are able to perform a synergistic degradation of rhamnogalacturonan. The deacetylation of the backbone by rhamnogalacturonan acetylesterase (RGAE) is an essential prerequisite for the subsequent action of the enzymes that cleave the glycosidic bonds. The structure of RGAE has been determined at 1.55 A resolution. RGAE folds into an alpha/beta/alpha structure. The active site of RGAE is an open cleft containing a serine-histidine-aspartic acid catalytic triad. The position of the three residues relative to the central parallel beta sheet and the lack of the nucleophilic elbow motif found in structures possessing the alpha/beta hydrolase fold show that RGAE does not belong to the alpha/beta hydrolase family. Structural and sequence comparisons have revealed that, despite very low sequence similarities, RGAE is related to seven other proteins. They are all members of a new hydrolase family, the SGNH-hydrolase family, which includes the carbohydrate esterase family 12 as a distinct subfamily. The SGNH-hydrolase family is characterised by having four conserved blocks of residues, each with one completely conserved residue; serine, glycine, asparagine and histidine, respectively. Each of the four residues plays a role in the catalytic function.
- Centre for Crystallographic Studies, University of Copenhagen, Copenhagen, DK-2100, Denmark.
Organizational Affiliation: