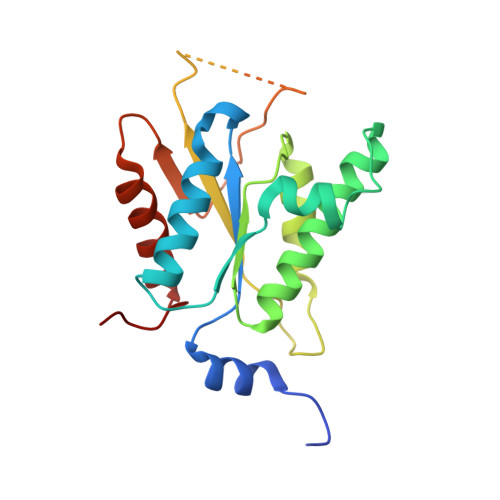
Crystal structure of adenosine 5'-phosphosulfate kinase from Penicillium chrysogenum.
MacRae, I.J., Segel, I.H., Fisher, A.J.(2000) Biochemistry 39: 1613-1621
- PubMed: 10677210 Search on PubMed
- DOI: https://doi.org/10.1021/bi9924157
- Primary Citation Related Structures:
1D6J - PubMed Abstract:
Adenosine 5'-phosphosulfate (APS) kinase catalyzes the second reaction in the two-step conversion of inorganic sulfate to 3'-phosphoadenosine 5'-phosphosulfate (PAPS). This report presents the 2.0 A resolution crystal structure of ligand-free APS kinase from the filamentous fungus, Penicillium chrysogenum. The enzyme crystallized as a homodimer with each subunit folded into a classic kinase motif consisting of a twisted, parallel beta-sheet sandwiched between two alpha-helical bundles. The Walker A motif, (32)GLSASGKS(39), formed the predicted P-loop structure. Superposition of the APS kinase active site region onto several other P-loop-containing proteins revealed that the conserved aspartate residue that usually interacts with the Mg(2+) coordination sphere of MgATP is absent in APS kinase. However, upon MgATP binding, a different aspartate, Asp 61, could shift and bind to the Mg(2+). The sequence (156)KAREGVIKEFT(166), which has been suggested to be a (P)APS motif, is located in a highly protease-susceptible loop that is disordered in both subunits of the free enzyme. MgATP or MgADP protects against proteolysis; APS alone has no effect but augments the protection provided by MgADP. The results suggest that the loop lacks a fixed structure until MgATP or MgADP is bound. The subsequent conformational change together with the potential change promoted by the interaction of MgATP with Asp 61 may define the APS binding site. This model is consistent with the obligatory ordered substrate binding sequence (MgATP or MgADP before APS) as established from steady state kinetics and equilibrium binding studies.
- Department of Chemistry and Section of Molecular and Cellular Biology, University of California, One Shields Avenue, Davis, California 95616, USA.
Organizational Affiliation: