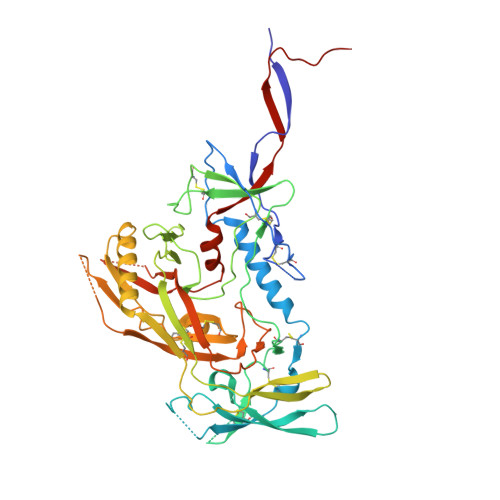
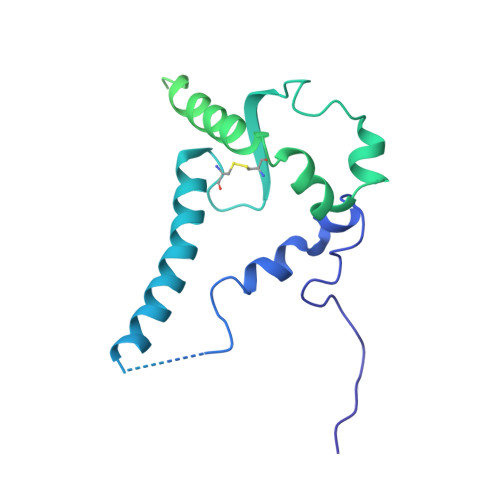
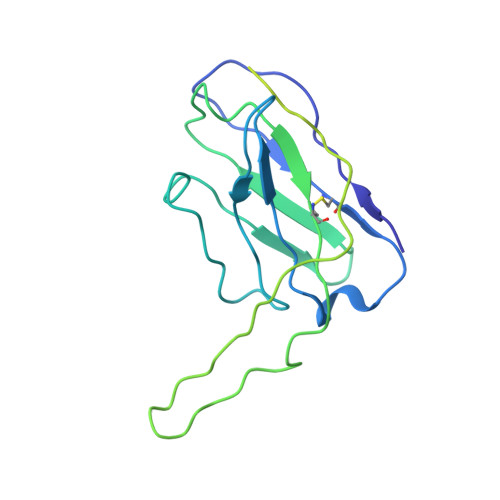
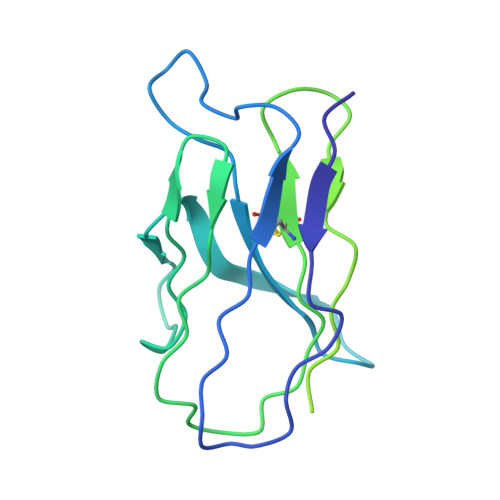
Co-evolution of HIV Envelope and Apex-Targeting Neutralizing Antibody Lineage Provides Benchmarks for Vaccine Design.
Rantalainen, K., Berndsen, Z.T., Murrell, S., Cao, L., Omorodion, O., Torres, J.L., Wu, M., Umotoy, J., Copps, J., Poignard, P., Landais, E., Paulson, J.C., Wilson, I.A., Ward, A.B.(2018) Cell Rep 23: 3249-3261
- PubMed: 29898396
- DOI: https://doi.org/10.1016/j.celrep.2018.05.046
- Primary Citation of Related Structures:
6CA6, 6CA7, 6CA9, 6DCQ - PubMed Abstract:
Broadly neutralizing antibodies (bnAbs) targeting the HIV envelope glycoprotein (Env) typically take years to develop. Longitudinal analyses of both neutralizing antibody lineages and viruses at serial time points during infection provide a basis for understanding the co-evolutionary contest between HIV and the humoral immune system. Here, we describe the structural characterization of an apex-targeting antibody lineage and autologous clade A viral Env from a donor in the Protocol C cohort. Comparison of Ab-Env complexes at early and late time points reveals that, within the antibody lineage, the CDRH3 loop rigidifies, the bnAb angle of approach steepens, and surface charges are mutated to accommodate glycan changes. Additionally, we observed differences in site-specific glycosylation between soluble and full-length Env constructs, which may be important for tuning optimal immunogenicity in soluble Env trimers. These studies therefore provide important guideposts for design of immunogens that prime and mature nAb responses to the Env V2-apex.
Organizational Affiliation:
Department of Integrative Structural and Computational Biology, The Scripps Research Institute, La Jolla, CA 92037, USA; IAVI Neutralizing Antibody Center and Collaboration of AIDS Vaccine Discovery, The Scripps Research Institute, La Jolla, CA 92037, USA.