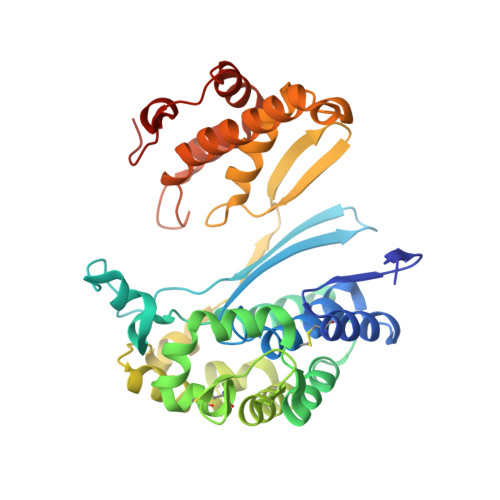
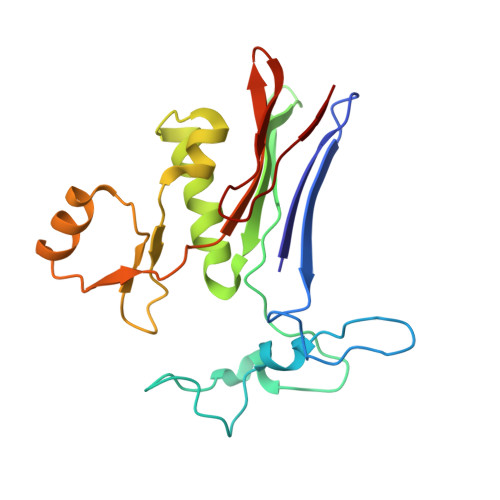
Structure of 6-diazo-5-oxo-norleucine-bound human gamma-glutamyl transpeptidase 1, a novel mechanism of inactivation.
Terzyan, S.S., Cook, P.F., Heroux, A., Hanigan, M.H.(2017) Protein Sci 26: 1196-1205
- PubMed: 28378915
- DOI: https://doi.org/10.1002/pro.3172
- Primary Citation of Related Structures:
5V4Q - PubMed Abstract:
Intense efforts are underway to identify inhibitors of the enzyme gamma-glutamyl transpeptidase 1 (GGT1) which cleaves extracellular gamma-glutamyl compounds and contributes to the pathology of asthma, reperfusion injury and cancer. The glutamate analog, 6-diazo-5-oxo-norleucine (DON), inhibits GGT1. DON also inhibits many essential glutamine metabolizing enzymes rendering it too toxic for use in the clinic as a GGT1 inhibitor. We investigated the molecular mechanism of human GGT1 (hGGT1) inhibition by DON to determine possible strategies for increasing its specificity for hGGT1. DON is an irreversible inhibitor of hGGT1. The second order rate constant of inactivation was 0.052 mM -1 min -1 and the K i was 2.7 ± 0.7 mM. The crystal structure of DON-inactivated hGGT1 contained a molecule of DON without the diazo-nitrogen atoms in the active site. The overall structure of the hGGT1-DON complex resembled the structure of the apo-enzyme; however, shifts were detected in the loop forming the oxyanion hole and elements of the main chain that form the entrance to the active site. The structure of hGGT1-DON complex revealed two covalent bonds between the enzyme and inhibitor which were part of a six membered ring. The ring included the OG atom of Thr381, the reactive nucleophile of hGGT1 and the α-amine of Thr381. The structure of DON-bound hGGT1 has led to the discovery of a new mechanism of inactivation by DON that differs from its inactivation of other glutamine metabolizing enzymes, and insight into the activation of the catalytic nucleophile that initiates the hGGT1 reaction.
Organizational Affiliation:
Laboratory of Biomolecular Structure and Function, University of Oklahoma Health Sciences Center, Oklahoma City, Oklahoma, 73104.