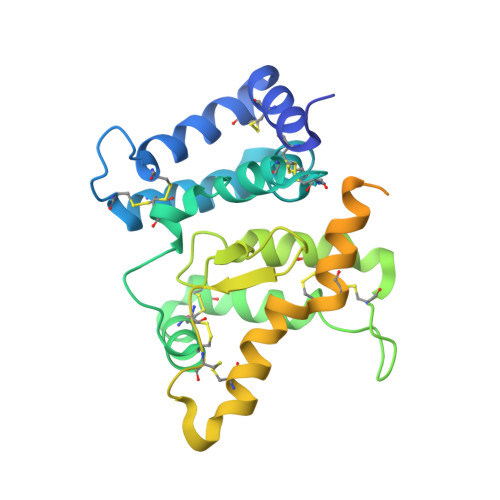
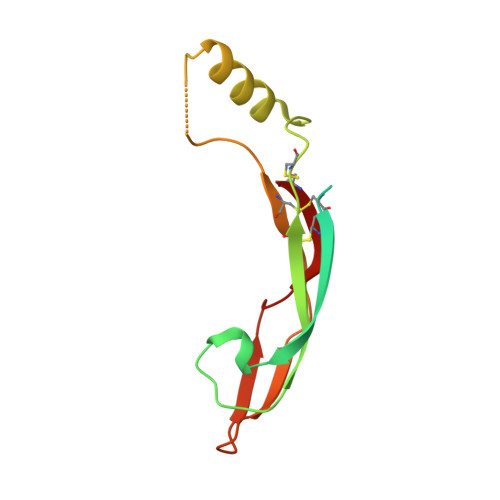
Comparison of GFL-GFRalpha complexes: further evidence relating GFL bend angle to RET signalling
Parkash, V., Goldman, A.(2009) Acta Crystallogr Sect F Struct Biol Cryst Commun 65: 551-558
- PubMed: 19478429
- DOI: https://doi.org/10.1107/S1744309109017722
- Primary Citation of Related Structures:
3FUB - PubMed Abstract:
Glial cell line-derived neurotrophic factor (GDNF) activates the receptor tyrosine kinase RET by binding to the GDNF-family receptor alpha1 (GFRalpha1) and forming the GDNF(2)-GFRalpha1(2)-RET(2) heterohexamer complex. A previous crystal structure of the GDNF(2)-GFRalpha1(2) complex (PDB code 2v5e) suggested that differences in signalling in GDNF-family ligand (GFL) complexes might arise from differences in the bend angle between the two monomers in the GFL homodimer. Here, a 2.35 A resolution structure of the GDNF(2)-GFRalpha1(2) complex crystallized with new cell dimensions is reported. The structure was refined to a final R factor of 22.5% (R(free) = 28%). The structures of both biological tetrameric complexes in the asymmetric unit are very similar to 2v5e and different from the artemin-GFRalpha3 structure, even though there is a small change in the structure of the GDNF. By comparison of all known GDNF and artemin structures, it is concluded that GDNF is more bent and more flexible than artemin and that this may be related to RET signalling. Comparisons also suggest that the differences between artemin and GDNF arise from the increased curvature of the artemin ;fingers', which both increases the buried surface area in the monomer-monomer interface and changes the intermonomer bend angle. From sequence comparison, it is suggested that neuturin (the second GFL) adopts an artemin-like conformation, while persephin has a different conformation to the other three.
Organizational Affiliation:
Institute of Biotechnology, University of Helsinki, Helsinki, Finland.