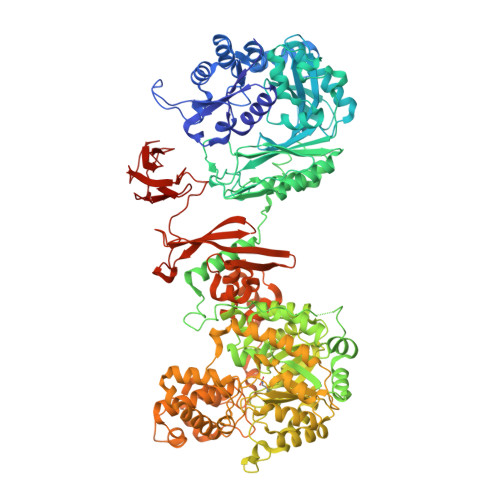
Domain architecture of pyruvate carboxylase, a biotin-dependent multifunctional enzyme
St Maurice, M., Reinhardt, L., Surinya, K.H., Attwood, P.V., Wallace, J.C., Cleland, W.W., Rayment, I.(2007) Science 317: 1076-1079
- PubMed: 17717183
- DOI: https://doi.org/10.1126/science.1144504
- Primary Citation of Related Structures:
2QF7 - PubMed Abstract:
Biotin-dependent multifunctional enzymes carry out metabolically important carboxyl group transfer reactions and are potential targets for the treatment of obesity and type 2 diabetes. These enzymes use a tethered biotin cofactor to carry an activated carboxyl group between distantly spaced active sites. The mechanism of this transfer has remained poorly understood. Here we report the complete structure of pyruvate carboxylase at 2.0 angstroms resolution, which shows its domain arrangement. The structure, when combined with mutagenic analysis, shows that intermediate transfer occurs between active sites on separate polypeptide chains. In addition, domain rearrangements associated with activator binding decrease the distance between active-site pairs, providing a mechanism for allosteric activation. This description provides insight into the function of biotin-dependent enzymes and presents a new paradigm for multifunctional enzyme catalysis.
Organizational Affiliation:
Department of Biochemistry, University of Wisconsin, Madison, WI 53706, USA.