Structural basis for viral late-domain binding to Alix
Lee, S., Joshi, A., Nagashima, K., Freed, E.O., Hurley, J.H.(2007) Nat Struct Mol Biol 14: 194-199
- PubMed: 17277784
- DOI: https://doi.org/10.1038/nsmb1203
- Primary Citation of Related Structures:
2OJQ - PubMed Abstract:
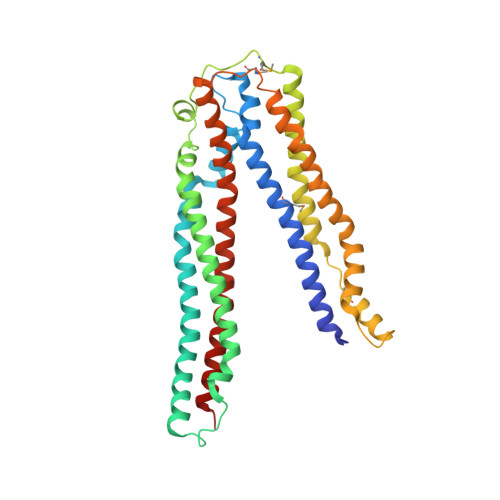
The modular protein Alix is a central node in endosomal-lysosomal trafficking and the budding of human immunodeficiency virus (HIV)-1. The Gag p6 protein of HIV-1 contains a LYPx(n)LxxL motif that is required for Alix-mediated budding and binds a region of Alix spanning residues 360-702. The structure of this fragment of Alix has the shape of the letter 'V' and is termed the V domain. The V domain has a topologically complex arrangement of 11 alpha-helices, with connecting loops that cross three times between the two arms of the V. The conserved residue Phe676 is at the center of a large hydrophobic pocket and is crucial for binding to a peptide model of HIV-1 p6. Overexpression of the V domain inhibits HIV-1 release from cells. This inhibition of release is reversed by mutations that block binding of the Alix V domain to p6.
Organizational Affiliation:
Laboratory of Molecular Biology, National Institute of Diabetes and Digestive and Kidney Diseases, National Institutes of Health (NIH), US Department of Health and Human Services, Bethesda, Maryland 20892, USA.