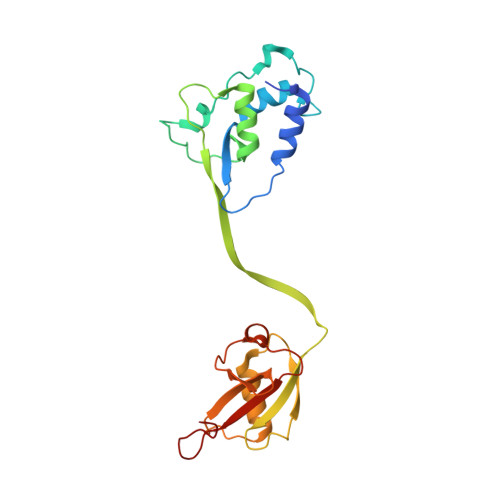
Structural variability of the ubiquitin specific protease DUSP-UBL double domains.
Elliott, P.R., Liu, H., Pastok, M.W., Grossmann, G.J., Rigden, D.J., Clague, M.J., Urbe, S., Barsukov, I.L.(2011) FEBS Lett 585: 3385-3390
- PubMed: 22001210
- DOI: https://doi.org/10.1016/j.febslet.2011.09.040
- Primary Citation of Related Structures:
3PV1, 4A3O, 4A3P - PubMed Abstract:
USP4, 11 and 15 are three closely related paralogues of the ubiquitin specific protease (USP) family of deubiquitinating enzymes. The DUSP domain and the UBL domain in these proteins are juxtaposed which may provide a functional unit conferring specificity. We determined the structures of the USP15 DUSP-UBL double domain unit in monomeric and dimeric states. We then conducted comparative analysis of the structural and physical properties of all three DUSP-UBL units. We identified structural features that dictate different dispositions between constituent domains, which in turn may influence respective binding properties.
Organizational Affiliation:
The University of Liverpool, Institute of Integrative Biology, Biosciences Building, Crown Street, Liverpool L69 7ZB, United Kingdom. P.R.Elliott@liv.ac.uk