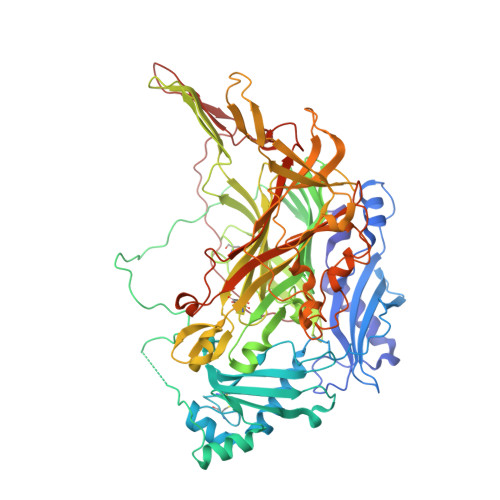
A new crystal form of human diamine oxidase.
McGrath, A.P., Hilmer, K.M., Collyer, C.A., Dooley, D.M., Guss, J.M.(2010) Acta Crystallogr Sect F Struct Biol Cryst Commun 66: 137-142
- PubMed: 20124708
- DOI: https://doi.org/10.1107/S1744309109052130
- Primary Citation of Related Structures:
3K5T - PubMed Abstract:
Copper amine oxidases (CAOs) are ubiquitous in nature and catalyse the oxidative deamination of primary amines to the corresponding aldehydes. Humans have three viable CAO genes (AOC1-3). AOC1 encodes human diamine oxidase (hDAO), which is the frontline enzyme for histamine metabolism. hDAO is unique among CAOs in that it has a distinct substrate preference for diamines. The structure of hDAO in space group P2(1)2(1)2(1) with two molecules in the asymmetric unit has recently been reported. Here, the structure of hDAO refined to 2.1 A resolution in space group C222(1) with one molecule in the asymmetric unit is reported.
Organizational Affiliation:
School of Molecular and Microbial Biosciences, University of Sydney, NSW 2006, Australia. a.mcgrath@mmb.usyd.edu.au