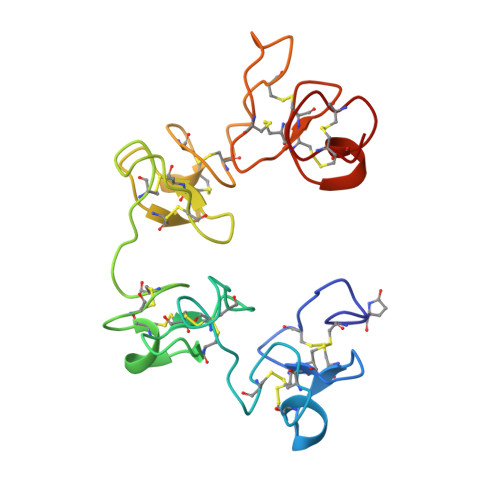
2.2 A resolution structure analysis of two refined N-acetylneuraminyl-lactose--wheat germ agglutinin isolectin complexes.
Wright, C.S.(1990) J Mol Biology 215: 635-651
- PubMed: 2231724 Search on PubMed
- DOI: https://doi.org/10.1016/S0022-2836(05)80174-3
- Primary Citation Related Structures:
1WGC, 2WGC, 7WGA, 9WGA - PubMed Abstract:
The crystal structures of complexes of isolectins 1 and 2 of wheat germ agglutinin (WGA1 and WGA2) with N-acetylneuraminyl-lactose (NeuNAc-alpha(2-3)-Gal-beta(1-4)-Glc) have been refined on the basis of data in the 8 to 2.2 A resolution range to final crystallographic R-factors of 17.2% and 15.3% (Fo greater than 1 sigma), respectively. Specific binding interactions and water association, as well as changes in conformation and mobility of the structure upon ligand binding, were compared in the two complexes. The temperature factors (B = 16.3 A2 and 18.4 A2) were found to be much lower compared with those of their respective native structures (19 to 22 A2). Residues involved in sugar binding, dimerization and in lattice contacts exhibit the largest decreases in B-value, suggesting that sugar binding reduces the overall mobility of the protein molecules in the crystal lattice. The binding mode of this sialyl-trisaccharide, an important cell receptor analogue, has been compared in the two isolectins. Only one of the two unique binding sites (4 per dimer), located in the subunit/subunit interface, is occupied in the crystals. This site, termed the "primary" binding site, contains one of the five amino acid substitutions that differentiate WGA1 and WGA2. Superposition of the refined models in each of the independent crystallographic environments indicates a close match only of the terminal non-reducing NeuNAc residue (root-mean-square delta r of 0.5 to 0.6 A). The Gal-Glc portion was found to superimpose poorly, lack electron density, and possess high atomic thermal factors. In both complexes NeuNAc is stabilized through contact with six amino acid side-chains (Ser114 and Glu115 of subunit 1 and Ser62, Tyr64, Tyr(His)66 and Tyr73 of subunit 2), involving all NeuNAc ring substituents. Refinement has allowed accurate assessment of the contact distances for four hydrogen bonds, a strong buried non-polar contact with the acetamido CH3 group and a large number of van der Waals' interactions with the three aromatic side-chains. The higher affinity of N-acetylneuraminyl-lactose observed by nuclear magnetic resonance studies for WGA1 can be explained by the more favorable binding interactions that occur when residue 66 is a Tyr. The tyrosyl side-chain provides a larger surface for van der Waals' stacking against the NeuNAc pyranose ring than His66 and a hydrogen bond contact with Gal (C2-OH), not possible in WGA2.(ABSTRACT TRUNCATED AT 400 WORDS)
- Department of Biochemistry and Molecular Biophysics, MCV/VCU, Richmond 23298-0001.
Organizational Affiliation: